Sandbox reserved 1753
From Proteopedia
(Difference between revisions)
| Line 75: | Line 75: | ||
== Introduction == | == Introduction == | ||
| - | Glycosylase is an enzyme. Its main function is in Base Excision Repair(BER) it removes and repairs damaged bases usually these are single stranded DNA breaks. BER corrects DNA damage that resulted from small leisures that do not disrupt the double helix. The way Glycosylase does this is by first cleaving the glycosidic bond of the damaged nucleotide leaving the Deoxyribose nucleotide with no base. The deoxyribose is then cleaved as well by AP endonuclease. The gap that is left is filled in through DNA Polymerase and DNA ligase. | + | Glycosylase is an enzyme. Its main function is in Base Excision Repair(BER) it removes and repairs damaged bases usually these are single stranded DNA breaks. BER corrects DNA damage that resulted from small leisures that do not disrupt the double helix. The way Glycosylase does this is by first cleaving the glycosidic bond of the damaged nucleotide leaving the Deoxyribose nucleotide with no base. The deoxyribose is then cleaved as well by AP endonuclease. The gap that is left is filled in through DNA Polymerase and DNA ligase. <scene name='92/927197/Active_site/1'>Active Site</scene> |
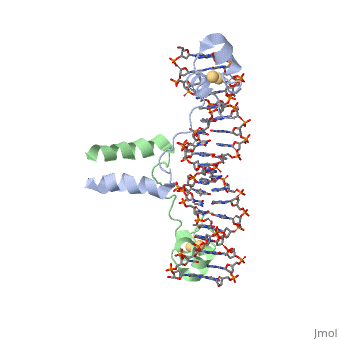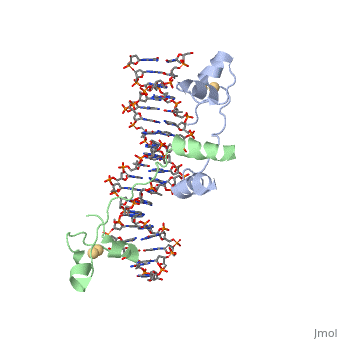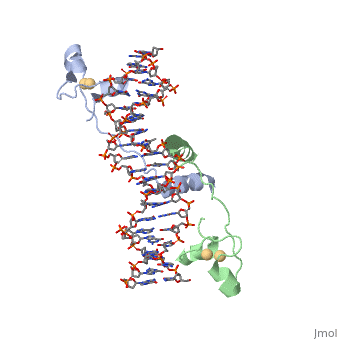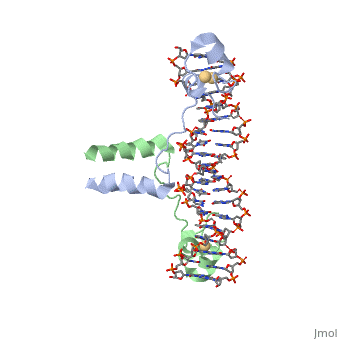
The structure of Glycosylase has a couple different forms in terms of its general structure. It is composed of a 10bp DNA that contains U G base pair mismatch. This is what allows it to bind the DNA flipping them out of the double helix. When the uracil mismatch is flipped out of the helix an <scene name='92/927197/Arg_side_chain/1'>ARG side chain</scene> takes its place. The actual structure is composed of a protein section bound to a DNA section. This is often represented by showing the DNA section in stick form and coloring it based on the different atoms bound. The protein section is characterized using a ribbon diagram. In our biochemistry book page 897 there is a representation of this as well. | The structure of Glycosylase has a couple different forms in terms of its general structure. It is composed of a 10bp DNA that contains U G base pair mismatch. This is what allows it to bind the DNA flipping them out of the double helix. When the uracil mismatch is flipped out of the helix an <scene name='92/927197/Arg_side_chain/1'>ARG side chain</scene> takes its place. The actual structure is composed of a protein section bound to a DNA section. This is often represented by showing the DNA section in stick form and coloring it based on the different atoms bound. The protein section is characterized using a ribbon diagram. In our biochemistry book page 897 there is a representation of this as well. | ||
Revision as of 13:06, 4 October 2022
==DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX==
| |||||||||||
A NUCLEOTIDE-FLIPPING MECHANISM FROM THE STRUCTURE OF HUMAN URACIL-DNA GLYCOSYLASE BOUND TO DNA
| |||||||||||
Categories: Atcc 18824 | Large Structures | Carey, M | Harrison, S C | Marmorstein, R | Ptashne, M | Double helix | Protein-dna complex | Transcription-dna complex | Human | Uridine nucleosidase | Arvai, A S | Kavli, B | Krokan, H E | Mol, C D | Slupphaug, G | Tainer, J A | Dna | Dna base excision repair | Dna glycosylase | Hydrolase-dna complex | Uracil