This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1757
From Proteopedia
(Difference between revisions)
| Line 14: | Line 14: | ||
Mevalonate 3,5-bisphosphate is a homodimer made up of <scene name='93/934001/Secondary/1'>40% alpha helices and 60% beta-sheets</scene>. Because Mevalonate 3,5-bisphosphate is a homodimer both molecules present are <scene name='93/934001/Quat/1'>identical</scene> in primary, secondary, and tertiary structure. Mevalonate 3,5-bisphosphate is amphipathic as showcased by the surplus of hydrophobic as well as hydrophilic amino acids present in its <scene name='93/934001/Active_site/2'>Active Site</scene>. This<scene name='93/934001/Active_site/4'> view</scene> also shows us the large cleft for substrate binding present on the molecule. | Mevalonate 3,5-bisphosphate is a homodimer made up of <scene name='93/934001/Secondary/1'>40% alpha helices and 60% beta-sheets</scene>. Because Mevalonate 3,5-bisphosphate is a homodimer both molecules present are <scene name='93/934001/Quat/1'>identical</scene> in primary, secondary, and tertiary structure. Mevalonate 3,5-bisphosphate is amphipathic as showcased by the surplus of hydrophobic as well as hydrophilic amino acids present in its <scene name='93/934001/Active_site/2'>Active Site</scene>. This<scene name='93/934001/Active_site/4'> view</scene> also shows us the large cleft for substrate binding present on the molecule. | ||
== Other important features == | == Other important features == | ||
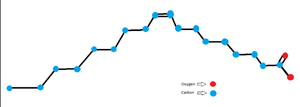
| + | As stated earlier the important amino acids section unfortunately the dimer for Mevalonate 3,5-bisphosphate has not been confirmed but one of the factors that led to the assumption that the unknown ligand is a fatty acid like Oleic Acid was due to the amphipathic nature of the binding site showcased by the presence of non-polar molecules in the binding site as well as polar molecules such as <scene name='93/934001/Binding_site/1'>Lys94, Tyr99, Arg128, and Glu138</scene> surrounding the opening of the cavity. | ||
</StructureSection> | </StructureSection> | ||
Revision as of 17:15, 13 December 2022
| This Sandbox is Reserved from November 4, 2022 through January 1, 2023 for use in the course CHEM 351 Biochemistry taught by Bonnie Hall at the Grand View University, Des Moines, USA. This reservation includes Sandbox Reserved 1755 through Sandbox Reserved 1764. |
To get started:
More help: Help:Editing |
Mevalonate 3,5-bisphosphate decarboxylase
| |||||||||||
References
[1]
- ↑ Aoki, M., Vinokur, J., Motoyama, K., Ishikawa, R., Collazo, M., Cascio, D., Sawaya, M. R., Ito, T., Bowie, J. U., & Hemmi, H. (2022). Crystal structure of mevalonate 3,5-bisphosphate decarboxylase reveals insight into the evolution of decarboxylases in the mevalonate metabolic pathways. The Journal of Biological Chemistry, 298(7), 102111–102111. https://doi.org/10.1016/j.jbc.2022.102111