This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1793
From Proteopedia
(Difference between revisions)
| Line 22: | Line 22: | ||
==== Bile Salt ==== | ==== Bile Salt ==== | ||
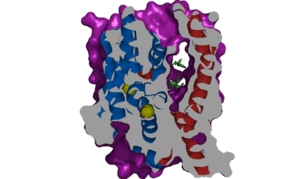
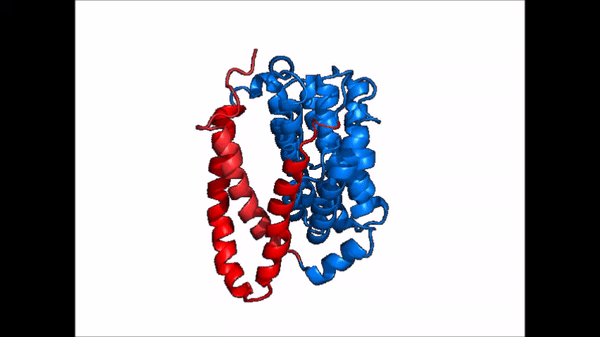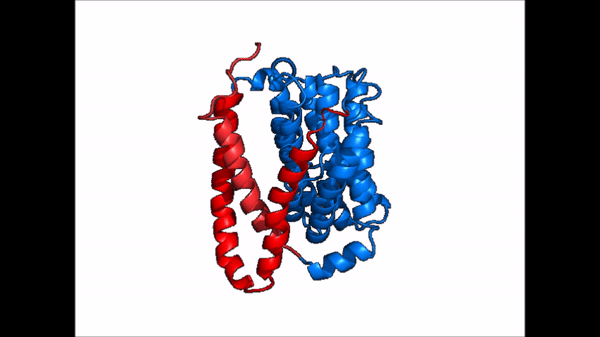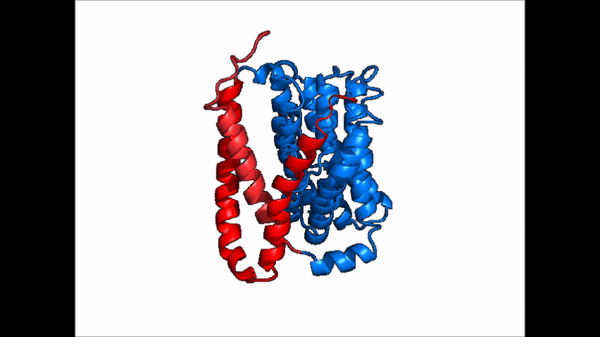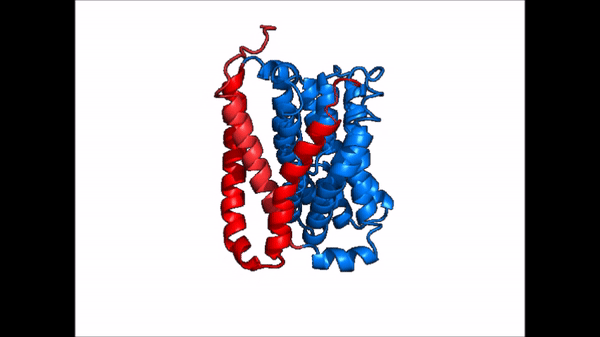
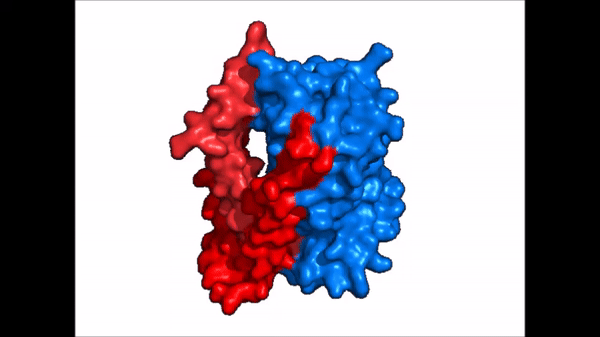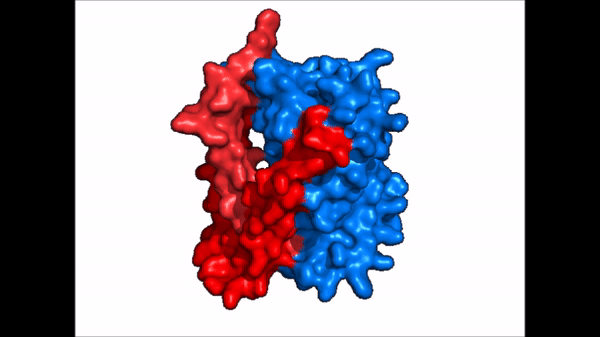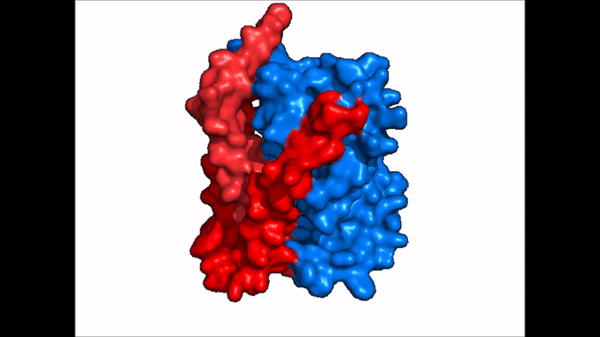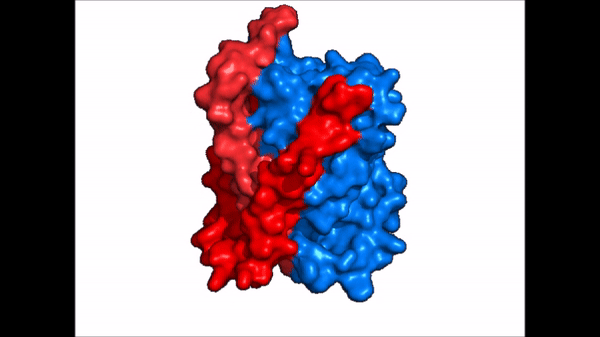
| - | The <scene name='95/952721/Amphipathic_patterns/ | + | The <scene name='95/952721/Amphipathic_patterns/2'>amphipathic pore</scene> is also characteristic of NTCP. The pore surface remains {{Template:ColorKey_Hydrophobic}}, while lining of the open pore state is largely {{Template:ColorKey_Polar}}. However, in the "inward-facing conformation" the polar pore residues are inaccessible. When the pore is closed only the surface hydrophobic residues are observed. As the pore opens up inner polar residues become accessible allowing for the binding of substrates. The pattern of hydrophobic and polar residues within the pore is believed to follow similar amphipathic patterns within taurocholate and other NTCP substrates, such as [https://en.wikipedia.org/wiki/Steroid steroids] and [https://en.wikipedia.org/wiki/Thyroid_hormones thyroid hormones]. <Ref name = Qi> Qi X. and Li W. (2022). Unlocking the secrets to human NTCP structure. The Innovation 3(5), 100294. https://doi.org/10.1016/j.xinn.2022.100294 </ref> Thus the channel provides specificity while preventing leakage of other substrates. When observing the relevant <scene name='95/952722/Bile_salts_res/1'>bile salt binding residues</scene> it is shown that some residues form Van der Waals interactions while others will form dipole-dipole or ionic interactions with bile salt substrates. The core domain appears to contribute most of the polar domains, while the panel domain contributes more hydrophobic residues. |
| - | + | ||
=== Conformational Change === | === Conformational Change === | ||
Revision as of 15:58, 6 April 2023
| This Sandbox is Reserved from February 27 through August 31, 2023 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1765 through Sandbox Reserved 1795. |
To get started:
More help: Help:Editing |
Sodium Bile Salt Co-Transporting Protein
| |||||||||||
References
- ↑ Stieger B. The role of the sodium-taurocholate cotransporting polypeptide (NTCP) and of the bile salt export pump (BSEP) in physiology and pathophysiology of bile formation. Handb Exp Pharmacol. 2011;(201):205-59. doi: 10.1007/978-3-642-14541-4_5. PMID: 21103971. DOI: DOI: 10.1007/978-3-642-14541-4_5.
- ↑ Anwer MS, Stieger B. Sodium-dependent bile salt transporters of the SLC10A transporter family: more than solute transporters. Pflugers Arch. 2014 Jan;466(1):77-89. PMID:24196564 doi:10.1007/s00424-013-1367-0
- ↑ Park, JH., Iwamoto, M., Yun, JH. et al. Structural insights into the HBV receptor and bile acid transporter NTCP. Nature 606, 1027–1031 (2022). https://doi.org/10.1038/s41586-022-04857-0.
- ↑ 4.0 4.1 Goutam, K., Ielasi, F.S., Pardon, E. et al. Structural basis of sodium-dependent bile salt uptake into the liver. Nature 606, 1015–1020 (2022). DOI: 10.1038/s41586-022-04723-z.
- ↑ Qi X. and Li W. (2022). Unlocking the secrets to human NTCP structure. The Innovation 3(5), 100294. https://doi.org/10.1016/j.xinn.2022.100294
- ↑ 6.0 6.1 Liu, H., Irobalieva, R.N., Bang-Sørensen, R. et al. Structure of human NTCP reveals the basis of recognition and sodium-driven transport of bile salts into the liver. Cell Res 32, 773–776 (2022). https://doi.org/10.1038/s41422-022-00680-4