This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1785
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | {{Template:CH462_Biochemistry_II_2023}}<!-- PLEASE ADD YOUR CONTENT BELOW HERE --> | ||
='''Human B-cell Antigen Receptor: IgM BCR'''= | ='''Human B-cell Antigen Receptor: IgM BCR'''= | ||
| - | <StructureSection load='7xQ8' size='350' side='right' caption='IgM B-Cell Receptor (PDB: 7xq8)' scene='95/952714/ | + | <StructureSection load='7xQ8' size='350' side='right' caption='IgM B-Cell Receptor (PDB: 7xq8)' scene='95/952714/Colored_by_chain/4'> |
=='''Introduction'''== | =='''Introduction'''== | ||
| - | ===Significance and Background=== | ||
Immunoglobulin M, or IgM, is one of multiple types of immunoglobulins that exist in humans. IgM presents itself on the surface of a B cell to act as a B Cell Receptor (BCR). Upon the binding of an antigen to the BCR, the B cell will activate, proliferate, and produce other Ig compounds. These include IgG, IgD, IgA, and IgE [https://proteopedia.org/wiki/index.php/Antibody antibodies], which all have specific roles in the various forms of immune response. Because of this, activation of the IgM BCR is a critical step in the beginning of an immune response. | Immunoglobulin M, or IgM, is one of multiple types of immunoglobulins that exist in humans. IgM presents itself on the surface of a B cell to act as a B Cell Receptor (BCR). Upon the binding of an antigen to the BCR, the B cell will activate, proliferate, and produce other Ig compounds. These include IgG, IgD, IgA, and IgE [https://proteopedia.org/wiki/index.php/Antibody antibodies], which all have specific roles in the various forms of immune response. Because of this, activation of the IgM BCR is a critical step in the beginning of an immune response. | ||
The structure of IgM was determined using [https://www.thermofisher.com/us/en/home/electron-microscopy/life-sciences/protein-analysis.html Cryo-EM] to visualize the atoms within the protein. However, due to poor resolution within specific regions of the IgM BCR, not every atom has been able to be visualized. | The structure of IgM was determined using [https://www.thermofisher.com/us/en/home/electron-microscopy/life-sciences/protein-analysis.html Cryo-EM] to visualize the atoms within the protein. However, due to poor resolution within specific regions of the IgM BCR, not every atom has been able to be visualized. | ||
| - | |||
| - | ===History and Discovery=== | ||
=='''Structure'''== | =='''Structure'''== | ||
| - | [[Image:Receptor_diagram.png|400 px|left|thumb|'''Figure 1. Cartoon Representation of IgM Antibody Receptor.''']] | ||
| - | <scene name='95/952714/Colored_by_domain/1'>colored by domain</scene> | ||
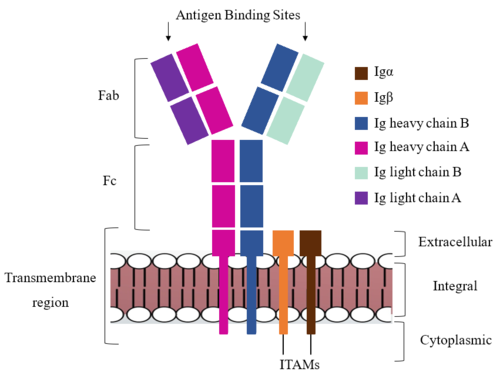
| - | <scene name='95/952713/Colored_by_chain/2'>colored by chain</scene> | + | The IgM BCR consists of six separate chains (Figure 1) that make up three main domains in the molecule. A depiction of the IgM <scene name='95/952714/Colored_by_domain/3'>colored by domain</scene> shows two heavy and two light chains together form the <b><span class="text-lightblue">Fab region</span></b>, or variable fragment at the top of the molecule where the antigen binding sites are located. The two heavy chains extend below the <b><span class="text-lightblue">Fab region</span></b> through the <b><span class="text-purple">Fc region</span></b> and eventually connect to the Igα/β heterodimer to form the <b><span class="text-orange">transmembrane region</span></b> which anchors the overall complex to the B cell. The overall structure, expression, and function of the IgM BCR has been found to be strongly influenced by the <b><span class="text-orange">transmembrane region</span></b> in which Ig α/β interactions as a heterodimer influence cell surface expression, receptor assembly, and effective signal transduction. In each domain, interactions between individual chains are important to understand the complex as a whole. All future 3D depictions will be <scene name='95/952713/Colored_by_chain/2'>colored by chain</scene> as in Figure 1. |
| + | [[Image:IgM_structure_overview_diagram.png|500 px|left|thumb|'''Figure 1. IgM BCR Structure Overview.''' Depiction of the IgM BCR expressed on the membrane of a B cell. Includes all major components including the α/β heterodimer, heavy and light chains, antigen binding sites, and the ITAM region for signal transduction.]] | ||
| + | {{Clear}} | ||
===Transmembrane Region=== | ===Transmembrane Region=== | ||
| - | The IgM BCR is anchored to [https://en.wikipedia.org/wiki/B_cell B-cell] membranes through the <scene name='95/952714/Integral_region/11'>transmembrane region</scene> which is broken up into both extracellular and integral domains which sit on top of or span through the membrane, respectively. IgM BCR assembly requires dimerization of the <b><span class="text-brown"> | + | The IgM BCR is anchored to [https://en.wikipedia.org/wiki/B_cell B-cell] membranes through the <scene name='95/952714/Integral_region/11'>transmembrane region</scene> which is broken up into both extracellular and integral domains which sit on top of or span through the membrane, respectively. IgM BCR assembly requires dimerization of the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> subunits which embed within the B-cell membrane. The <scene name='95/952714/Ig_alpha_beta/5'>Igα and Igβ heterodimer</scene> dimerizes within the extracellular region with a <scene name='95/952714/Extracellular_disulfide_bridge/6'>disulfide bridge</scene>. Additional dimerization is believed to occur within the integral region via a hydrogen bond; the involved residues and interaction have not been confirmed. Although the mechanism of disulfide bridge formation is still unknown, it is believed that <scene name='95/952714/Extracellular_glycosylation/2'>extracellular glycosylation</scene> via <b><span class="text-lightgreen">N-linked glycosylation</span></b> (NAGs) on various asparagine residues in the extracellular region of both the <b><span class="text-brown">Igα</span></b> and and <b><span class="text-orange">Igβ</span></b> chains help facilitate this process. [https://en.wikipedia.org/wiki/Chaperone_(protein) Chaperone proteins] remain bound to the alpha and beta subunits until both dimerizations occur; at this point the rest of the BCR complex can be recruited. |
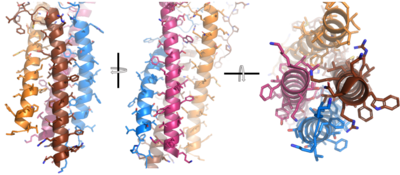
| - | + | After <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> dimerization, the transmembrane helices of the heavy chains can embed within the B-cell membrane. The side chains of this <scene name='95/952714/Integral_helices_2/2'>4-pass integral helix structure</scene> are primarily hydrophobic side chains that allow for interactions with the hydrophobic tails in the [https://en.wikipedia.org/wiki/Lipid_bilayer phospholipid bilayer]. The 4 helices (Figure 2) are primarily held together through hydrophobic interactions; however, a a few polar residues are included on the interior of the helix structure which interact with a few polar residues on the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> chains. | |
| - | + | Furthermore, both the Igα and Igβ chains have cytoplasmic tails that extend into the B cell (Figure 1). Each of these tails contain a immuno-receptor tyrosine-based activation motif (ITAM) region to facilitate signal transduction (Figure 4). | |
| + | [[Image:Integral_helix_figure.png|400 px|left|thumb|'''Figure 2. 4-pass integral helix.''' Pymol image of the integral helices in IgM BCR (PDB:7xq8) rotated on the x and y axes. Side chains are shown as sticks. Brown=Ig alpha, orange=Ig beta, pink=heavy chain A, blue=heavy chain B.]] | ||
| + | {{Clear}} | ||
===Fc Region=== | ===Fc Region=== | ||
| + | |||
The constant region of IgM is made up of the 2 <scene name='95/952715/Heavy_chain/1'>heavy chains</scene>. These heavy chains form a bridge to connect the Fab fragment, or variable region, to the transmembrane region. They also act as a wire to allow the variable region to send a cellular signal through to the intermembrane region once an antigen has been bound. | The constant region of IgM is made up of the 2 <scene name='95/952715/Heavy_chain/1'>heavy chains</scene>. These heavy chains form a bridge to connect the Fab fragment, or variable region, to the transmembrane region. They also act as a wire to allow the variable region to send a cellular signal through to the intermembrane region once an antigen has been bound. | ||
| Line 33: | Line 32: | ||
<scene name='95/952713/Disulfides/4'>heavy chain disulfide bond</scene> | <scene name='95/952713/Disulfides/4'>heavy chain disulfide bond</scene> | ||
| + | |||
| + | <scene name='95/952713/Trans_heavy/5'>heavy chain A heavy chain B interface</scene> | ||
<scene name='95/952713/Trans_heavy/2'>transmembrane heavy chain interface</scene> | <scene name='95/952713/Trans_heavy/2'>transmembrane heavy chain interface</scene> | ||
===Fab Region=== | ===Fab Region=== | ||
| - | Because the Fab region of IgM is so poorly resolved, a structural comparison to another antibody was performed to approximate where an antigen would bind to the <scene name='95/952713/Variable_region/1'>variable region</scene>. Figure 1 on the left shows the antigen binding motif (located on the Fab region) of an Ig-BCR complex that was engineered to contain the variable region of a neutralizing antibody called VCR01, an antibody that targets the epitope of HIV molecules. It contains areas referred to as complementary-determining regions, or CDRs, which are where the antigen makes contact with the antibody on the Fab domain. Showing them as surface representation allows us to make structural comparisons to the IgM antibody, and to highlight their similarities. The CDRs are similarly placed within the heavy and light chain variable regions between both antibodies. It is speculated that they are structurally similar because the VCR01 antibody can effectively target multiple HIV strains while IgM is the preliminary antibody produced and released during early stages of the immune response, thus it is able to respond in larger concentrations while antibodies that are more specific to the antigen are being produced. | ||
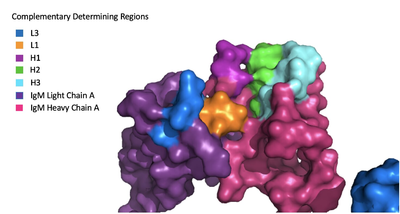
| - | + | Because the Fab region of IgM is so poorly resolved, a structural comparison to another antibody was performed to approximate where an antigen would bind to the <scene name='95/952713/Variable_region/1'>variable region</scene>. Figure 2 shows the antigen binding motif (located on the Fab region) of an Ig-BCR complex that was engineered to contain the variable region of a neutralizing antibody called VCR01, an antibody that targets the epitope of HIV molecules. It contains areas referred to as complementary-determining regions, or CDRs, which are where the antigen makes contact with the antibody on the Fab domain. Showing them as surface representation allows us to make structural comparisons to the IgM antibody, and to highlight their similarities. The CDRs are similarly placed within the heavy and light chain variable regions between both antibodies. It is speculated that they are structurally similar because the VCR01 antibody can effectively target multiple HIV strains while IgM is the preliminary antibody produced and released during early stages of the immune response, thus it is able to respond in larger concentrations while antibodies that are more specific to the antigen are being produced. | |
Due to the Fab region of the IgM antibody being poorly resolved, the specific side chain interactions between the heavy and light chains have not been determined. This depiction of the <scene name='95/952713/Heavy-light_chain_interface/1'>heavy-light chain interface</scene> shows how the 4 β-sandwiches fit together; heavy chain A and B of the Fab region form a complex with the rest of the molecule via interactions with the heavy A and B of the Fc region, before continuing down into the intracellular domain to interact with the transmembrane region. The light chains however are only connected to the complex by forming interactions with the heavy chains within the Fab region. Although the specific residues within the Fab region have yet to be identified, it is estimated that each β-sandwich contains one disulfide bridge with additional hydrogen bonds. | Due to the Fab region of the IgM antibody being poorly resolved, the specific side chain interactions between the heavy and light chains have not been determined. This depiction of the <scene name='95/952713/Heavy-light_chain_interface/1'>heavy-light chain interface</scene> shows how the 4 β-sandwiches fit together; heavy chain A and B of the Fab region form a complex with the rest of the molecule via interactions with the heavy A and B of the Fc region, before continuing down into the intracellular domain to interact with the transmembrane region. The light chains however are only connected to the complex by forming interactions with the heavy chains within the Fab region. Although the specific residues within the Fab region have yet to be identified, it is estimated that each β-sandwich contains one disulfide bridge with additional hydrogen bonds. | ||
| - | + | [[Image:Igm_surface.png|400 px|left|thumb|'''Figure 3. Surface Representation of IgM Antibody Binding Pocket.''']] | |
| - | == | + | {{Clear}} |
| - | + | ||
| + | =='''Signal Transduction'''== | ||
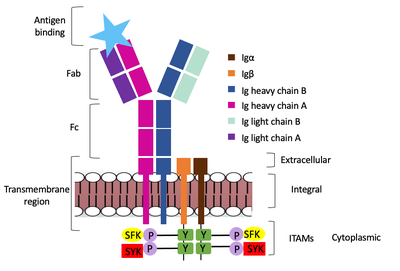
| - | + | The diagram in Figure 4 depicts the initial process of B cell activation by the antigen binding to the antibody at the Fab region. The underlying mechanism for signal transduction is unknown but it is speculated to operate under what is known as the conserved assembly mechanism. This means that upon antigen binding, BCRs on the surface of the cell begin to cluster to cause the phosphorylation of the immunoreceptor tyrosine-based activation motifs located in Igα and Igβ. In its “off” state, the constant region 4 of heavy chain B overlaps the extracellular components of Igα and Igβ. As the antigen binds, it induces a conformational change to release the overlap and allow for clustering about the BCR. Now, in its “on” state the phosphorylation of the ITAM region (observed here as the conserved tyrosine residues are phosphorylated) within the intracellular tails of Igα and Igβ drives downstream kinase activity to continue to process of signal cascading. | |
| + | [[Image:Signal_transduction-2.png|400 px|left|thumb|'''Figure 4. IgM Antibody Signal Transduction following Antigen Binding.''']] | ||
| + | {{Clear}} | ||
</StructureSection> | </StructureSection> | ||
Revision as of 01:12, 7 April 2023
Human B-cell Antigen Receptor: IgM BCR
| |||||||||||
References
Student Contributors
Detonyeá Dickson, Allison Goss, Jackson Payton