This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1806
From Proteopedia
(Difference between revisions)
| Line 25: | Line 25: | ||
== Structural highlights == | == Structural highlights == | ||
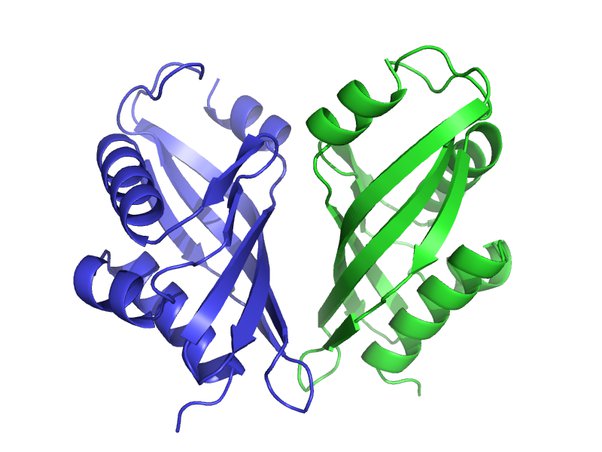
| - | The secondary structure of a protein contains regions of amino acid chains that are stabilized by hydrogen bonds which create alpha-helix and beta-pleated sheets. For Pyrrolysine tRNA synthetase, the secondary structure has 2 domains, one called the N-terminal and the other being the C-terminal. Pyrrolysine is a modified lysine with a 4-methylpyrroline-5-carboxylate group linked by an amide to the ɛ-amino group. Pyrrolysine is lysine in which has a pyrroline ring linked to the end of the lysine side chain. | + | The secondary structure of a protein contains regions of amino acid chains that are stabilized by hydrogen bonds which create alpha-helix and beta-pleated sheets. For Pyrrolysine tRNA synthetase, the <scene name='95/954103/Secondary_structure/1'>secondary structure</scene> has 2 domains, one called the N-terminal and the other being the C-terminal. Pyrrolysine is a modified lysine with a 4-methylpyrroline-5-carboxylate group linked by an amide to the ɛ-amino group. Pyrrolysine is lysine in which has a pyrroline ring linked to the end of the lysine side chain. |
The ''tertiary structure'' of a protein is the overall 3D structure of the protein. The protein is stabilized by outside polar hydrophilic hydrogen and ionic bond interactions, and internal hydrophobic interactions between non-polar amino acid side chains. | The ''tertiary structure'' of a protein is the overall 3D structure of the protein. The protein is stabilized by outside polar hydrophilic hydrogen and ionic bond interactions, and internal hydrophobic interactions between non-polar amino acid side chains. | ||
Revision as of 22:37, 27 April 2023
| This Sandbox is Reserved from Mar 1 through Jun 1, 2023 for use in the course CHEM 351 Biochemistry taught by Bonnie_Hall at the Grand View University, Des Moines, USA. This reservation includes Sandbox Reserved 1796 through Sandbox Reserved 1811. |
To get started:
More help: Help:Editing |
Pyrrolysine (2Q7H) Structure
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Kavran JM, Gundllapalli S, O'Donoghue P, Englert M, Soll D, Steitz TA. Structure of pyrrolysyl-tRNA synthetase, an archaeal enzyme for genetic code innovation. Proc Natl Acad Sci U S A. 2007 Jul 3;104(27):11268-73. Epub 2007 Jun 25. PMID:17592110
- ↑ Nozawa K, O'Donoghue P, Gundllapalli S, Araiso Y, Ishitani R, Umehara T, Soll D, Nureki O. Pyrrolysyl-tRNA synthetase-tRNA(Pyl) structure reveals the molecular basis of orthogonality. Nature. 2009 Feb 26;457(7233):1163-7. Epub 2008 Dec 31. PMID:19118381 doi:10.1038/nature07611