This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
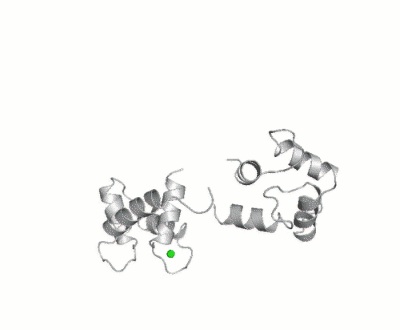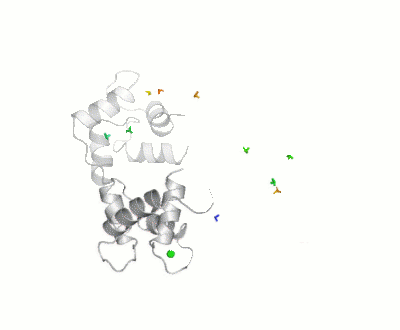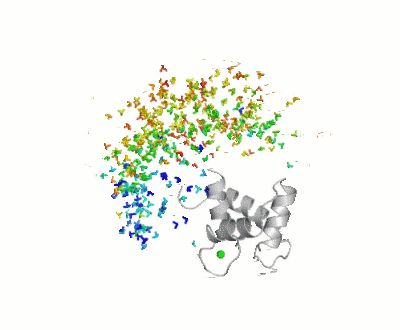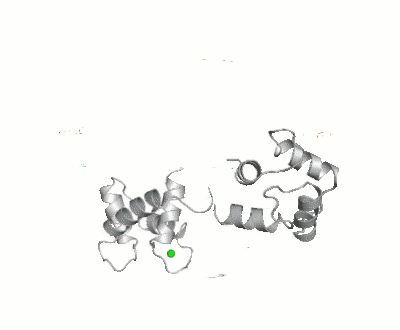
Calmodulin
From Proteopedia
(Difference between revisions)
| Line 20: | Line 20: | ||
<jmolButton> | <jmolButton> | ||
<script> | <script> | ||
| - | script "/wiki/images/ | + | script "/wiki/images/a/a2/Storymorph.spt"; |
load files "=1prw" "=1cll"; | load files "=1prw" "=1cll"; | ||
delete water; | delete water; | ||
| Line 41: | Line 41: | ||
structures = [{1.1}, {2.1}]; | structures = [{1.1}, {2.1}]; | ||
my_recipe = [ | my_recipe = [ | ||
| - | [ | + | [{79-147},{79-147}, {(79-138) and alpha}], |
| - | [ | + | [{4-78}, {79}, {(8-78) and alpha}], |
]; | ]; | ||
| - | morph(20, structures, my_recipe | + | morph(20, structures, my_recipe); |
</script> | </script> | ||
<text>Open</text> | <text>Open</text> | ||
| Line 57: | Line 57: | ||
structures = [{2.1}, {1.1}]; | structures = [{2.1}, {1.1}]; | ||
my_recipe = [ | my_recipe = [ | ||
| - | [ | + | [{79-147},{79-147}, {(79-138) and alpha}], |
| - | [ | + | [{4-78}, {79}, {(8-78) and alpha}], |
]; | ]; | ||
| - | morph(20, structures, my_recipe | + | morph(20, structures, my_recipe); |
</script> | </script> | ||
<text>Close</text> | <text>Close</text> | ||
| Line 70: | Line 70: | ||
<jmolButton> | <jmolButton> | ||
<script> | <script> | ||
| - | script "/wiki/images/ | + | script "/wiki/images/a/a2/Storymorph.spt"; |
load files "=1prw" "=1cll"; | load files "=1prw" "=1cll"; | ||
delete water; | delete water; | ||
| Line 91: | Line 91: | ||
structures = [{1.1}, {2.1}]; | structures = [{1.1}, {2.1}]; | ||
my_recipe = [ | my_recipe = [ | ||
| - | [ | + | [{79-147},{79-147}, {(79-138) and alpha}], |
| - | [ | + | [{4-78}, {79}, {(8-78) and alpha}], |
]; | ]; | ||
| - | morph(20, structures, my_recipe | + | morph(20, structures, my_recipe); |
</script> | </script> | ||
<text>Open</text> | <text>Open</text> | ||
| Line 107: | Line 107: | ||
structures = [{2.1}, {1.1}]; | structures = [{2.1}, {1.1}]; | ||
my_recipe = [ | my_recipe = [ | ||
| - | [ | + | [{79-147},{79-147}, {(79-138) and alpha}], |
| - | [ | + | [{4-78}, {79}, {(8-78) and alpha}], |
]; | ]; | ||
| - | morph(20, structures, my_recipe | + | morph(20, structures, my_recipe); |
</script> | </script> | ||
<text>Close</text> | <text>Close</text> | ||
| Line 120: | Line 120: | ||
<jmolButton> | <jmolButton> | ||
<script> | <script> | ||
| - | script "/wiki/images/ | + | script "/wiki/images/a/a2/Storymorph.spt"; |
load files "=1prw" "=1cll"; | load files "=1prw" "=1cll"; | ||
delete water; | delete water; | ||
| Line 141: | Line 141: | ||
structures = [{1.1}, {2.1}]; | structures = [{1.1}, {2.1}]; | ||
my_recipe = [ | my_recipe = [ | ||
| - | [ | + | [{4-78}, {4-78}, {(8-78) and alpha}], |
| - | [ | + | [{79-147},{78}, {(79-138) and alpha}], |
]; | ]; | ||
| - | morph(20, structures, my_recipe | + | morph(20, structures, my_recipe); |
</script> | </script> | ||
<text>Open</text> | <text>Open</text> | ||
| Line 157: | Line 157: | ||
structures = [{2.1}, {1.1}]; | structures = [{2.1}, {1.1}]; | ||
my_recipe = [ | my_recipe = [ | ||
| - | [ | + | [{4-78}, {4-78}, {(8-78) and alpha}], |
| - | [ | + | [{79-147},{78}, {(79-138) and alpha}], |
]; | ]; | ||
| - | morph(20, structures, my_recipe | + | morph(20, structures, my_recipe); |
</script> | </script> | ||
<text>Close</text> | <text>Close</text> | ||
Current revision
| |||||||||||
See Also
Bibliography
- ↑ Chattopadhyaya R, Meador WE, Means AR, Quiocho FA. Calmodulin structure refined at 1.7 A resolution. J Mol Biol. 1992 Dec 20;228(4):1177-92. PMID:1474585
- ↑ Bertini I, Giachetti A, Luchinat C, Parigi G, Petoukhov MV, Pierattelli R, Ravera E, Svergun DI. Conformational Space of Flexible Biological Macromolecules from Average Data. J Am Chem Soc. 2010 Sep 7. PMID:20822180 doi:10.1021/ja1063923
- ↑ The Storymorph Jmol scripts creates the interpolated coordinates of the morph on the fly.
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Karsten Theis, Jaime Prilusky, Enrico Ravera, Daniel Moyano-Marino