This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1bg1
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='1bg1' size='340' side='right'caption='[[1bg1]], [[Resolution|resolution]] 2.25Å' scene=''> | <StructureSection load='1bg1' size='340' side='right'caption='[[1bg1]], [[Resolution|resolution]] 2.25Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1bg1]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/ | + | <table><tr><td colspan='2'>[[1bg1]] is a 2 chain structure with sequence from [https://en.wikipedia.org/wiki/Mus_musculus Mus musculus]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1BG1 OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1BG1 FirstGlance]. <br> |
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.25Å</td></tr> |
| + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=PTR:O-PHOSPHOTYROSINE'>PTR</scene></td></tr> | ||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1bg1 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1bg1 OCA], [https://pdbe.org/1bg1 PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1bg1 RCSB], [https://www.ebi.ac.uk/pdbsum/1bg1 PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1bg1 ProSAT]</span></td></tr> | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1bg1 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1bg1 OCA], [https://pdbe.org/1bg1 PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1bg1 RCSB], [https://www.ebi.ac.uk/pdbsum/1bg1 PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1bg1 ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
| - | + | [https://www.uniprot.org/uniprot/STAT3_MOUSE STAT3_MOUSE] Signal transducer and transcription activator that mediates cellular responses to interleukins, KITLG/SCF and other growth factors. May mediate cellular responses to activated FGFR1, FGFR2, FGFR3 and FGFR4. Binds to the interleukin-6 (IL-6)-responsive elements identified in the promoters of various acute-phase protein genes. Activated by IL31 through IL31RA. STAT3B interacts with the N-terminal part of JUN to activate such promoters in a cooperative way.<ref>PMID:11294897</ref> | |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 19: | Line 20: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1bg1 ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1bg1 ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
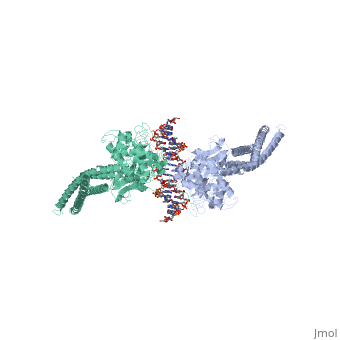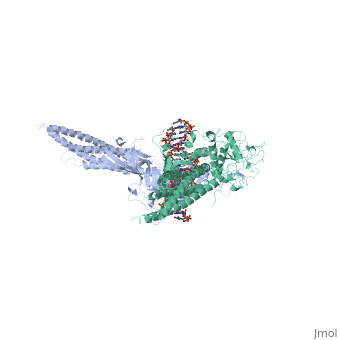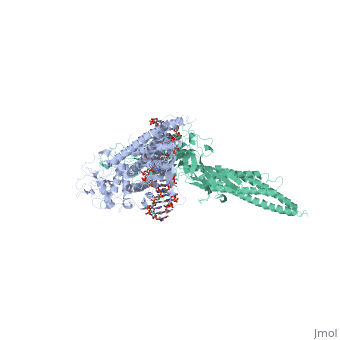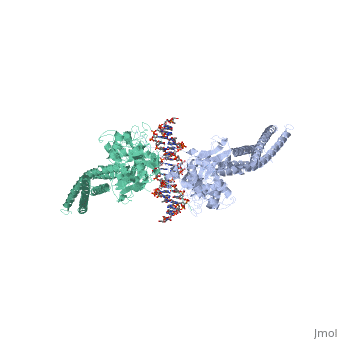
| - | STAT proteins are a family of eukaryotic transcription factors that mediate the response to a large number of cytokines and growth factors. Upon activation by cell-surface receptors or their associated kinases, STAT proteins dimerize, translocate to the nucleus and bind to specific promoter sequences on their target genes. Here we report the first crystal structure of a STAT protein bound to its DNA recognition site at 2.25 A resolution. The structure provides insight into the various steps by which STAT proteins deliver a response signal directly from the cell membrane to their target genes in the nucleus. | ||
| - | |||
| - | Three-dimensional structure of the Stat3beta homodimer bound to DNA.,Becker S, Groner B, Muller CW Nature. 1998 Jul 9;394(6689):145-51. PMID:9671298<ref>PMID:9671298</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 1bg1" style="background-color:#fffaf0;"></div> | ||
== References == | == References == | ||
<references/> | <references/> | ||
| Line 33: | Line 25: | ||
</StructureSection> | </StructureSection> | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: | + | [[Category: Mus musculus]] |
| - | [[Category: Becker | + | [[Category: Becker S]] |
| - | [[Category: Groner | + | [[Category: Groner B]] |
| - | [[Category: Muller | + | [[Category: Muller CW]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
TRANSCRIPTION FACTOR STAT3B/DNA COMPLEX
| |||||||||||