This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1nfk
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
== Structural highlights == | == Structural highlights == | ||
<table><tr><td colspan='2'>[[1nfk]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Mus_musculus Mus musculus]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1NFK OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1NFK FirstGlance]. <br> | <table><tr><td colspan='2'>[[1nfk]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Mus_musculus Mus musculus]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1NFK OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1NFK FirstGlance]. <br> | ||
| - | </td></tr><tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1nfk FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1nfk OCA], [https://pdbe.org/1nfk PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1nfk RCSB], [https://www.ebi.ac.uk/pdbsum/1nfk PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1nfk ProSAT]</span></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.3Å</td></tr> |
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1nfk FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1nfk OCA], [https://pdbe.org/1nfk PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1nfk RCSB], [https://www.ebi.ac.uk/pdbsum/1nfk PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1nfk ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
| - | + | [https://www.uniprot.org/uniprot/NFKB1_MOUSE NFKB1_MOUSE] NF-kappa-B is a pleiotropic transcription factor present in almost all cell types and is the endpoint of a series of signal transduction events that are initiated by a vast array of stimuli related to many biological processes such as inflammation, immunity, differentiation, cell growth, tumorigenesis and apoptosis. NF-kappa-B is a homo- or heterodimeric complex formed by the Rel-like domain-containing proteins RELA/p65, RELB, NFKB1/p105, NFKB1/p50, REL and NFKB2/p52 and the heterodimeric p65-p50 complex appears to be most abundant one. The dimers bind at kappa-B sites in the DNA of their target genes and the individual dimers have distinct preferences for different kappa-B sites that they can bind with distinguishable affinity and specificity. Different dimer combinations act as transcriptional activators or repressors, respectively. NF-kappa-B is controlled by various mechanisms of post-translational modification and subcellular compartmentalization as well as by interactions with other cofactors or corepressors. NF-kappa-B complexes are held in the cytoplasm in an inactive state complexed with members of the NF-kappa-B inhibitor (I-kappa-B) family. In a conventional activation pathway, I-kappa-B is phosphorylated by I-kappa-B kinases (IKKs) in response to different activators, subsequently degraded thus liberating the active NF-kappa-B complex which translocates to the nucleus. NF-kappa-B heterodimeric p65-p50 and RelB-p50 complexes are transcriptional activators. The NF-kappa-B p50-p50 homodimer is a transcriptional repressor, but can act as a transcriptional activator when associated with BCL3. NFKB1 appears to have dual functions such as cytoplasmic retention of attached NF-kappa-B proteins by p105 and generation of p50 by a cotranslational processing. The proteasome-mediated process ensures the production of both p50 and p105 and preserves their independent function, although processing of NFKB1/p105 also appears to occur post-translationally. p50 binds to the kappa-B consensus sequence 5'-GGRNNYYCC-3', located in the enhancer region of genes involved in immune response and acute phase reactions. Plays a role in the regulation of apoptosis. Isoform 5, isoform 6 and isoform 7 act as inhibitors of transactivation of p50 NF-kappa-B subunit, probably by sequestering it in the cytoplasm. Isoform 3 (p98) (but not p84 or p105) acts as a transactivator of NF-kappa-B-regulated gene expression. In a complex with MAP3K8, NFKB1/p105 represses MAP3K8-induced MAPK signaling; active MAP3K8 is released by proteasome-dependent degradation of NFKB1/p105. | |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 18: | Line 19: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1nfk ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1nfk ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
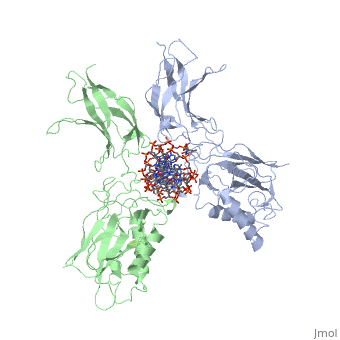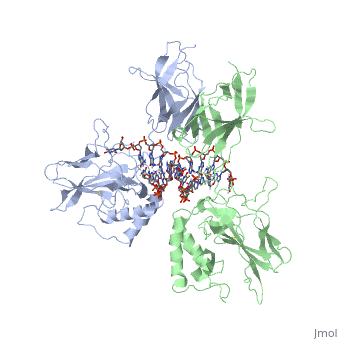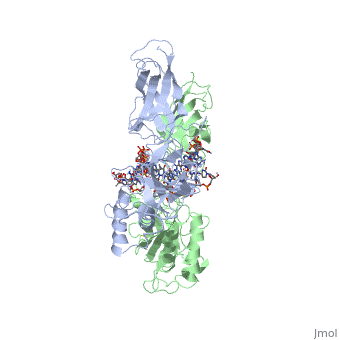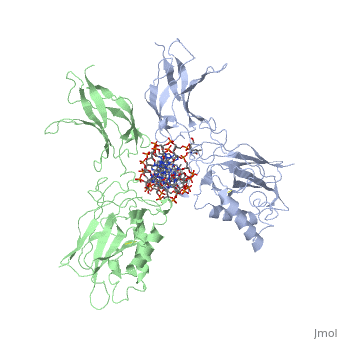
| - | The 2.3-A crystal structure of the transcription factor NK-kappa B p50 homodimer bound to a palindromic kappa B site reveals that the Rel homology region folds into two distinct domains, similar to those in the immunoglobulin superfamily. The p50 dimer envelopes an undistorted B-DNA helix, making specific contacts along the 10-base-pair kappa B recognition site mainly through loops connecting secondary structure elements in both domains. The carboxy-terminal domains form a dimerization interface between beta-sheets using residues that are strongly conserved in the Rel family. | ||
| - | |||
| - | Structure of NF-kappa B p50 homodimer bound to a kappa B site.,Ghosh G, van Duyne G, Ghosh S, Sigler PB Nature. 1995 Jan 26;373(6512):303-10. PMID:7530332<ref>PMID:7530332</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 1nfk" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
*[[NF-kB|NF-kB]] | *[[NF-kB|NF-kB]] | ||
| - | == References == | ||
| - | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
[[Category: Mus musculus]] | [[Category: Mus musculus]] | ||
| - | + | [[Category: Ghosh G]] | |
| - | [[Category: Ghosh | + | [[Category: Ghosh S]] |
| - | [[Category: Ghosh | + | [[Category: Sigler PB]] |
| - | [[Category: Sigler | + | [[Category: Van Duyne G]] |
| - | [[Category: | + | |
| - | + | ||
Current revision
STRUCTURE OF THE NUCLEAR FACTOR KAPPA-B (NF-KB) P50 HOMODIMER
| |||||||||||