We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Andrew Helmerich Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 13: | Line 13: | ||
<scene name='10/1038819/Amylin_pep2/5'>Amylin Overview</scene> | <scene name='10/1038819/Amylin_pep2/5'>Amylin Overview</scene> | ||
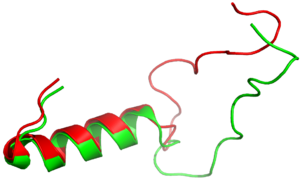
| - | [[Image: | + | [[Image:align.png|300px|left|thumb|Figure Legend]] |
== Structure == | == Structure == | ||
Revision as of 13:22, 18 April 2024
The Amylin Receptor(AMYR)
| |||||||||||
References
- ↑ Ransey E, Paredes E, Dey SK, Das SR, Heroux A, Macbeth MR. Crystal structure of the Entamoeba histolytica RNA lariat debranching enzyme EhDbr1 reveals a catalytic Zn(2+) /Mn(2+) heterobinucleation. FEBS Lett. 2017 Jul;591(13):2003-2010. doi: 10.1002/1873-3468.12677. Epub 2017, Jun 14. PMID:28504306 doi:http://dx.doi.org/10.1002/1873-3468.12677
- ↑ Cao J, Belousoff MJ, Liang YL, Johnson RM, Josephs TM, Fletcher MM, Christopoulos A, Hay DL, Danev R, Wootten D, Sexton PM. A structural basis for amylin receptor phenotype. Science. 2022 Mar 25;375(6587):eabm9609. PMID:35324283 doi:10.1126/science.abm9609
Student Contributors
- Ty Vander Eide
- Andrew Helmerich
- Ben Whiteside