This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Isabel Kluszynski/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 24: | Line 24: | ||
===Binding Interactions of Tirzepatide=== | ===Binding Interactions of Tirzepatide=== | ||
| - | Tirzepatide is an imbalanced GLP-1R/GIP-R coagonist. Tirzepatide has an equal affinity for the GIP-R as native GIP does, but it has a lower affinity for GLP-1R than native GLP-1 does. <ref name= | + | Tirzepatide is an imbalanced GLP-1R/GIP-R coagonist. Tirzepatide has an equal affinity for the GIP-R as native GIP does, but it has a lower affinity for GLP-1R than native GLP-1 does. <ref name="Sun">PMID:35333651</ref> Because of this, there is a biased signaling that results from Tirzepatide binding. This leads to greater cAMP generation and lower beta-arrestin recruitment resulting in a lesser degree of GLP-1R internalization. This means more GLP-1R is exposed on the surface of cells. |
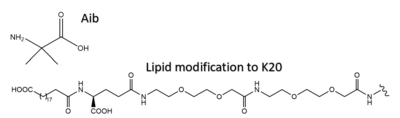
[[Image:Aib_and_C20.png|400 px|right|thumb|Structure of 2-Aminoisobutyric acid and the C20 fatty diacid moiety. Tirzepatide is modified with Aib at position 2 and 13 and the fatty diacid at K20.]] | [[Image:Aib_and_C20.png|400 px|right|thumb|Structure of 2-Aminoisobutyric acid and the C20 fatty diacid moiety. Tirzepatide is modified with Aib at position 2 and 13 and the fatty diacid at K20.]] | ||
| - | Several residues have been selectively and intentionally modified during Tirzepatide drug design. <scene name='10/1037487/Interesting_tirz_modifications/3'>Aib</scene>, aminoisobutyric acid, is located at position 2 and 13 with the purpose of preventing DPP-4 cleavage. K20 of Tirzepatide has a lipid modification, specifically a C20 fatty diacid moiety, that serves to enhance binding to the protein carrier albumin and increase the half-life of the drug in the body. <ref name= | + | Several residues have been selectively and intentionally modified during Tirzepatide drug design. <scene name='10/1037487/Interesting_tirz_modifications/3'>Aib</scene>, aminoisobutyric acid, is located at position 2 and 13 with the purpose of preventing DPP-4 cleavage. K20 of Tirzepatide has a lipid modification, specifically a C20 fatty diacid moiety, that serves to enhance binding to the protein carrier albumin and increase the half-life of the drug in the body. <ref name="Sun"/> |
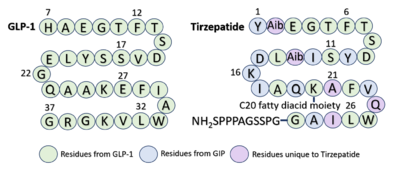
Similarly to GLP-1 forming several stabilizing interactions with GLP-1R, Tirzepatide also forms many key <scene name='10/1037490/Tirzepatidebonding/9'>stabilizing interactions</scene>. Beginning at the N terminus of Tirzepatide (Tirz), <scene name='10/1037490/Tirzepatidebonding/11'>Tirz E3</scene> forms a salt bridge with GLP-1R R190 and a hydrogen bond with GLP-1R Y152. Additionally, Tirz T7 hydrogen bonds with GLP-1R K197. These interactions are nearly identical to the interactions GLP-1 E9 and T13 make with the receptor. Towards the middle of the peptide, <scene name='10/1037490/Tirzepatidebonding/10'>Tirz D15</scene> forms a hydrogen bond with Y205, yet another similar interaction to GLP-1 binding. <ref name="Zhao"/> Looking at the sequence comparison of GLP-1 and Tirzepatide, GLP-1 E21 is located at the same position as Tirz D15. | Similarly to GLP-1 forming several stabilizing interactions with GLP-1R, Tirzepatide also forms many key <scene name='10/1037490/Tirzepatidebonding/9'>stabilizing interactions</scene>. Beginning at the N terminus of Tirzepatide (Tirz), <scene name='10/1037490/Tirzepatidebonding/11'>Tirz E3</scene> forms a salt bridge with GLP-1R R190 and a hydrogen bond with GLP-1R Y152. Additionally, Tirz T7 hydrogen bonds with GLP-1R K197. These interactions are nearly identical to the interactions GLP-1 E9 and T13 make with the receptor. Towards the middle of the peptide, <scene name='10/1037490/Tirzepatidebonding/10'>Tirz D15</scene> forms a hydrogen bond with Y205, yet another similar interaction to GLP-1 binding. <ref name="Zhao"/> Looking at the sequence comparison of GLP-1 and Tirzepatide, GLP-1 E21 is located at the same position as Tirz D15. | ||
| Line 36: | Line 36: | ||
===Comparison of Binding Interactions=== | ===Comparison of Binding Interactions=== | ||
| - | Through an <scene name='10/1037490/Overlay/7'>overlay</scene>, it is visible that GLP-1 and Tirzepatide bind similarly but not identically to the receptor. Some interactions seen in GLP-1 bound to GLP-1R are preserved with Tirzepatide, while others are altered. For example, the first residue of GLP-1 is a <scene name='10/1037487/Gvt_binding/6'>histidine</scene>, while the first residue of Tirzepatide is a <scene name='10/1037490/Gvt_binding/11'>tyrosine</scene>. <ref name= | + | Through an <scene name='10/1037490/Overlay/7'>overlay</scene>, it is visible that GLP-1 and Tirzepatide bind similarly but not identically to the receptor. Some interactions seen in GLP-1 bound to GLP-1R are preserved with Tirzepatide, while others are altered. For example, the first residue of GLP-1 is a <scene name='10/1037487/Gvt_binding/6'>histidine</scene>, while the first residue of Tirzepatide is a <scene name='10/1037490/Gvt_binding/11'>tyrosine</scene>. <ref name="Sun"/> The identity of this first residue can either favor or prevent specific GLP-1R interactions. With GLP-1 H7, <scene name='10/1037490/Gvt_binding/8'>GLP-1R R310</scene> <scene name='10/1037487/Gvt_binding/4'>GLP-1R R310</scene> and E373 are in close enough proximity to form a salt bridge. Additionally, GLP-1R W306 and D372 are positioned ideally for hydrogen bond formation. However, with Tirzepatide bound, Tirz Y1 causes greater steric clashing, pushing the <scene name='10/1037490/Gvt_binding/9'>GLP-1R R310</scene> <scene name='10/1037487/Gvt_binding/5'>GLP-1R R310</scene> and other residues further apart so they are unable to interact. <ref name="Sun"/> |
| - | Another example of a difference in receptor confirmation can be seen with GLP-1R R299. With GLP-1 bound, <scene name='10/1037487/Glp1_comparison/2'>GLP-1R R299</scene> faces the peptide and is able to hydrogen bond with either GLP-1 S17 or E21. When Tirzepatide is bound, <scene name='10/1037487/Tirzepatide/6'>GLP-1R R299</scene> flips away from the peptide and can no longer hydrogen bond with any Tirzepatide residues. <ref name= | + | Another example of a difference in receptor confirmation can be seen with GLP-1R R299. With GLP-1 bound, <scene name='10/1037487/Glp1_comparison/2'>GLP-1R R299</scene> faces the peptide and is able to hydrogen bond with either GLP-1 S17 or E21. When Tirzepatide is bound, <scene name='10/1037487/Tirzepatide/6'>GLP-1R R299</scene> flips away from the peptide and can no longer hydrogen bond with any Tirzepatide residues. <ref name="Sun"/> Additionally, differential binding of GLP-1 and Tirzepatide modifies how the transmembrane domain interacts with the G-alpha subunit to initiate a signal cascade. For example, when GLP-1 is bound to the receptor, <scene name='10/1037487/G_alpha_glp/2'>GLP-1R F257</scene> is able to participate in a pi stacking interaction with G-alpha F376. When Tirzepatide is bound, <scene name='10/1037487/G_alpha_tirz/3'>GLP-1R 257</scene> is oriented facing away from the G-alpha subunit. <ref name="Zhao"/> |
Referencing the sequence alignment, GLP-1 F28 and W31 are in the same position as Tirzepatide F22 and W25, indicating that the phenylalanine and tryptophan residues are conserved among the two sequences. As discussed previously in the GLP-1 binding interactions subsection, <scene name='10/1037487/Cterm_glp/8'>GLP-1 F28</scene> and W31 participate in a pi stacking interaction with GLP-1R W214. Similarly, <scene name='10/1037487/Tirzepatide/5'>Tirz F22</scene> and W25 are also able to interact aromatically with GLP-1R W214. | Referencing the sequence alignment, GLP-1 F28 and W31 are in the same position as Tirzepatide F22 and W25, indicating that the phenylalanine and tryptophan residues are conserved among the two sequences. As discussed previously in the GLP-1 binding interactions subsection, <scene name='10/1037487/Cterm_glp/8'>GLP-1 F28</scene> and W31 participate in a pi stacking interaction with GLP-1R W214. Similarly, <scene name='10/1037487/Tirzepatide/5'>Tirz F22</scene> and W25 are also able to interact aromatically with GLP-1R W214. | ||
Revision as of 13:12, 25 April 2024
=GLP-1R Homo Sapiens=
| |||||||||||
References
- ↑ 1.0 1.1 1.2 Mayendraraj A, Rosenkilde MM, Gasbjerg LS. GLP-1 and GIP receptor signaling in beta cells interactions and co-stimulation. Peptides. 2022 May;151:170749. PMID:35065096 doi:10.1016/j.peptides.2022.170749
- ↑ 2.0 2.1 Seino Y, Fukushima M, Yabe D. GIP and GLP-1, the two incretin hormones: Similarities and differences. J Diabetes Investig. 2010 Apr 22;1(1-2):8-23. PMID:24843404 doi:10.1111/j.2040-1124.2010.00022.x
- ↑ Zhang X, Belousoff MJ, Zhao P, Kooistra AJ, Truong TT, Ang SY, Underwood CR, Egebjerg T, Šenel P, Stewart GD, Liang YL, Glukhova A, Venugopal H, Christopoulos A, Furness SGB, Miller LJ, Reedtz-Runge S, Langmead CJ, Gloriam DE, Danev R, Sexton PM, Wootten D. Differential GLP-1R Binding and Activation by Peptide and Non-peptide Agonists. Mol Cell. 2020 Nov 5;80(3):485-500.e7. PMID:33027691 doi:10.1016/j.molcel.2020.09.020
- ↑ 4.0 4.1 4.2 Zhao F, Zhou Q, Cong Z, Hang K, Zou X, Zhang C, Chen Y, Dai A, Liang A, Ming Q, Wang M, Chen LN, Xu P, Chang R, Feng W, Xia T, Zhang Y, Wu B, Yang D, Zhao L, Xu HE, Wang MW. Structural insights into multiplexed pharmacological actions of tirzepatide and peptide 20 at the GIP, GLP-1 or glucagon receptors. Nat Commun. 2022 Feb 25;13(1):1057. PMID:35217653 doi:10.1038/s41467-022-28683-0
- ↑ 5.0 5.1 5.2 5.3 5.4 Sun B, Willard FS, Feng D, Alsina-Fernandez J, Chen Q, Vieth M, Ho JD, Showalter AD, Stutsman C, Ding L, Suter TM, Dunbar JD, Carpenter JW, Mohammed FA, Aihara E, Brown RA, Bueno AB, Emmerson PJ, Moyers JS, Kobilka TS, Coghlan MP, Kobilka BK, Sloop KW. Structural determinants of dual incretin receptor agonism by tirzepatide. Proc Natl Acad Sci U S A. 2022 Mar 29;119(13):e2116506119. PMID:35333651 doi:10.1073/pnas.2116506119
Student Contributors
- Isabel Kluszynski
- Makenna Marcinek