User:Chloe Tucker/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 19: | Line 19: | ||
[[Image:GIP_hydrogen_bonds.jpg|350 px|right|thumb|Figure 1. GIPR and GIP residue interactions]] | [[Image:GIP_hydrogen_bonds.jpg|350 px|right|thumb|Figure 1. GIPR and GIP residue interactions]] | ||
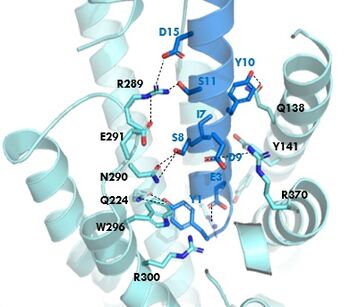
The <scene name='10/1038815/Overview/3'>binding site</scene> of GIP with the GIP receptor (GIPR) is where the N-term of GIP binds with the transmembrane domain of the GIPR. The first interaction formed with GIPR and the N-term of GIP is a hydrogen bond between Tyrosine 1 (Y1) and Glutamine 224 (Q224) to activate the G-protein to start sending signals to the cell. | The <scene name='10/1038815/Overview/3'>binding site</scene> of GIP with the GIP receptor (GIPR) is where the N-term of GIP binds with the transmembrane domain of the GIPR. The first interaction formed with GIPR and the N-term of GIP is a hydrogen bond between Tyrosine 1 (Y1) and Glutamine 224 (Q224) to activate the G-protein to start sending signals to the cell. | ||
| - | Many other <scene name='10/1038815/Active_site/3'>residues</scene> within the binding site are forming hydrogen bonds and hydrophobic interactions between the ligand and the receptor. The N-term binds more strongly than the C-term and there are many different residues contributing to this, including, Tyrosine 1 (Y1) and Tryptophan (W296) are forming aromatic interactions. Serine 8 (S8) and Asparagine 290 (N290) are forming two hydrogen bonds with each other. Aspartate 9 (D9) is forming another hydrogen bond with Arginine 370 (R370). These hydrogen bonds lead to the activation of cell signaling and when this binding is somehow disrupted, that is what causes different diseases like diabetes. | + | Many other <scene name='10/1038815/Active_site/3'>residues</scene> within the binding site are forming hydrogen bonds and hydrophobic interactions between the ligand and the receptor. The N-term binds more strongly than the C-term and there are many different residues contributing to this, including, Tyrosine 1 (Y1) and Tryptophan (W296) are forming aromatic interactions. Serine 8 (S8) and Asparagine 290 (N290) are forming two hydrogen bonds with each other. Aspartate 9 (D9) is forming another hydrogen bond with Arginine 370 (R370)<ref name=”Sun”/>. These hydrogen bonds lead to the activation of cell signaling and when this binding is somehow disrupted, that is what causes different diseases like diabetes. |
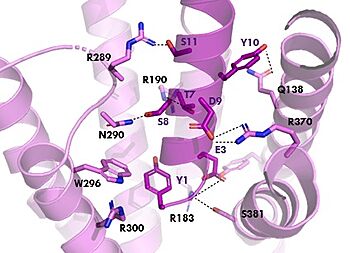
=== Binding/Active Site of GIPR with Tirzepatide === | === Binding/Active Site of GIPR with Tirzepatide === | ||
Revision as of 13:46, 25 April 2024
GIP and GIP-R
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Sun B, Willard FS, Feng D, Alsina-Fernandez J, Chen Q, Vieth M, Ho JD, Showalter AD, Stutsman C, Ding L, Suter TM, Dunbar JD, Carpenter JW, Mohammed FA, Aihara E, Brown RA, Bueno AB, Emmerson PJ, Moyers JS, Kobilka TS, Coghlan MP, Kobilka BK, Sloop KW. Structural determinants of dual incretin receptor agonism by tirzepatide. Proc Natl Acad Sci U S A. 2022 Mar 29;119(13):e2116506119. PMID:35333651 doi:10.1073/pnas.2116506119
Student Contributors
- Chloe Tucker
- Mandy Bechman