Kemp eliminase
From Proteopedia
(Difference between revisions)
| Line 1: | Line 1: | ||
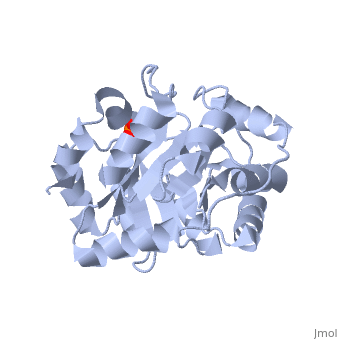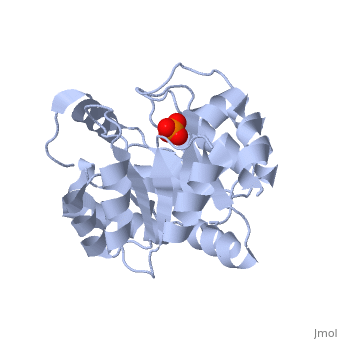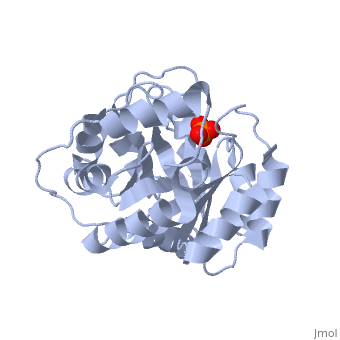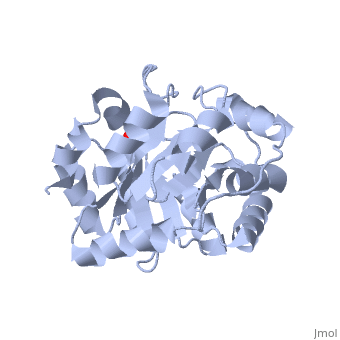
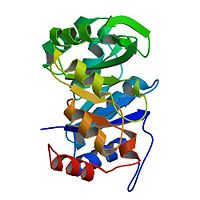
<StructureSection load='3uxa' size='400' scene='' caption='Designed kemp eliminase KE59 complex with phosphate (PDB code [[3uxa]])'> | <StructureSection load='3uxa' size='400' scene='' caption='Designed kemp eliminase KE59 complex with phosphate (PDB code [[3uxa]])'> | ||
[[Image:3iio.jpg|left|200px]] | [[Image:3iio.jpg|left|200px]] | ||
| + | __TOC__ | ||
{{ABSTRACT_PUBMED_20036254}} | {{ABSTRACT_PUBMED_20036254}} | ||
A series of [http://en.wikipedia.org/wiki/Protein_design computationally designed enzymes] that catalyze the Kemp elimination have described. Kemp eliminase (KE07) has <scene name='3iio/Int/10'>TIM barrel scaffold</scene>. The Kemp elimination of <scene name='3iio/Int/4'>5-nitrobenzisoxazole</scene> was chosen as a model reaction for [http://en.wikipedia.org/wiki/Proton#Hydrogen_as_proton proton (H)] transfer from [http://en.wikipedia.org/wiki/Carbon carbon], simultaneously with the cut of the [http://en.wikipedia.org/wiki/Nitrogen nitrogen]–[http://en.wikipedia.org/wiki/Oxygen oxygen] (N-O) bond, resulting in <scene name='3iio/Int/8'>cyanophenol product</scene>. Such reaction is a critical step in many [http://en.wikipedia.org/wiki/Enzyme enzymatic reactions]. The [http://en.wikipedia.org/wiki/Enzyme_catalysis catalytic] base (E101), the general acid/[http://en.wikipedia.org/wiki/Hydrogen_bond H-bond] donor (K222), and the stacking residue (W50) make interactions with the 5-nitrobenzisoxazole at the <scene name='3iio/Int/9'>active site of KE07</scene>. [http://en.wikipedia.org/wiki/Directed_evolution Directed evolution] can significantly improve the stability, expression and activity of enzymes. In the catalytically improved directed evolutionary variants of KE07 containing the <scene name='3iio/Ali/7'>Ile7Asp mutation</scene>, Asp7 breaks the Glu101–Lys222 salt bridge (for example [[3iiv]], chain A is shown). | A series of [http://en.wikipedia.org/wiki/Protein_design computationally designed enzymes] that catalyze the Kemp elimination have described. Kemp eliminase (KE07) has <scene name='3iio/Int/10'>TIM barrel scaffold</scene>. The Kemp elimination of <scene name='3iio/Int/4'>5-nitrobenzisoxazole</scene> was chosen as a model reaction for [http://en.wikipedia.org/wiki/Proton#Hydrogen_as_proton proton (H)] transfer from [http://en.wikipedia.org/wiki/Carbon carbon], simultaneously with the cut of the [http://en.wikipedia.org/wiki/Nitrogen nitrogen]–[http://en.wikipedia.org/wiki/Oxygen oxygen] (N-O) bond, resulting in <scene name='3iio/Int/8'>cyanophenol product</scene>. Such reaction is a critical step in many [http://en.wikipedia.org/wiki/Enzyme enzymatic reactions]. The [http://en.wikipedia.org/wiki/Enzyme_catalysis catalytic] base (E101), the general acid/[http://en.wikipedia.org/wiki/Hydrogen_bond H-bond] donor (K222), and the stacking residue (W50) make interactions with the 5-nitrobenzisoxazole at the <scene name='3iio/Int/9'>active site of KE07</scene>. [http://en.wikipedia.org/wiki/Directed_evolution Directed evolution] can significantly improve the stability, expression and activity of enzymes. In the catalytically improved directed evolutionary variants of KE07 containing the <scene name='3iio/Ali/7'>Ile7Asp mutation</scene>, Asp7 breaks the Glu101–Lys222 salt bridge (for example [[3iiv]], chain A is shown). | ||
Revision as of 12:29, 7 July 2024
| |||||||||||
See also
3D structures of Kemp eliminase
Updated on 07-July-2024
References
- Khersonsky O, Rothlisberger D, Dym O, Albeck S, Jackson CJ, Baker D, Tawfik DS. Evolutionary optimization of computationally designed enzymes: Kemp eliminases of the KE07 series. J Mol Biol. 2010 Mar 5;396(4):1025-42. Epub 2009 Dec 28. PMID:20036254 doi:10.1016/j.jmb.2009.12.031
- Rothlisberger D, Khersonsky O, Wollacott AM, Jiang L, DeChancie J, Betker J, Gallaher JL, Althoff EA, Zanghellini A, Dym O, Albeck S, Houk KN, Tawfik DS, Baker D. Kemp elimination catalysts by computational enzyme design. Nature. 2008 May 8;453(7192):190-5. Epub 2008 Mar 19. PMID:18354394 doi:10.1038/nature06879
- Khersonsky O, Kiss G, Rothlisberger D, Dym O, Albeck S, Houk KN, Baker D, Tawfik DS. Bridging the gaps in design methodologies by evolutionary optimization of the stability and proficiency of designed Kemp eliminase KE59. Proc Natl Acad Sci U S A. 2012 Jun 8. PMID:22685214 doi:10.1073/pnas.1121063109
- ↑ Labas A, Szabo E, Mones L, Fuxreiter M. Optimization of reorganization energy drives evolution of the designed Kemp eliminase KE07. Biochim Biophys Acta. 2013 May;1834(5):908-17. PMID:23380188 doi:10.1016/j.bbapap.2013.01.005
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, Jaime Prilusky
Categories: Escherichia coli | Dym, O. | Khersonsky, O. | Tawfik, D S. | Beta barrel | Lyase | Albeck, S. | ISPC, Israel Structural Proteomics Center. | Alpha-beta barrel | Amino-acid biosynthesis | Cytoplasm | Histidine biosynthesis | ISPC | Israel Structural Proteomics Center | Structural genomic | Topic Page