Electrostatic potential maps
From Proteopedia
(Difference between revisions)
(→Gallery) |
(→Gallery) |
||
| Line 18: | Line 18: | ||
|- | |- | ||
| [[Image:Electrostatic potential 1tsj.PNG|200 px]] || Electrostatic potential map of [[1tsj]] made with the [https://epmv.scripps.edu/ Embedded Python Molecular Viewer] from the [https://ccsb.scripps.edu/ Center for Computational Structural Biology] of the Scripps Research Institute. | | [[Image:Electrostatic potential 1tsj.PNG|200 px]] || Electrostatic potential map of [[1tsj]] made with the [https://epmv.scripps.edu/ Embedded Python Molecular Viewer] from the [https://ccsb.scripps.edu/ Center for Computational Structural Biology] of the Scripps Research Institute. | ||
| - | |} | ||
Click on the image to enlarge. | Click on the image to enlarge. | ||
Revision as of 18:16, 25 August 2024
It is revealing to visualize the distribution of electrostatic charges, electrostatic potential, on molecular surfaces. Most protein-protein and protein-ligand interactions are largely electrostatic in nature, via hydrogen bonds and ionic interactions. Their strengths are modulated by the nature of the solvent: pure water or high ionic strength aqueous solution.
Gallery
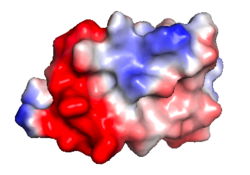
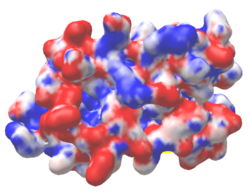
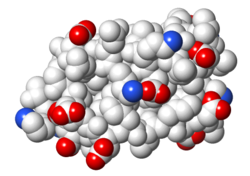
| Protein 1pgb is in the same orientation in all images. Positive + / Negative - | ||
|---|---|---|
| 
| 
|
| Electrostatic potential map rendered by PyMOL. | Electrostatic potential map rendered by iCn3D. | Van der Waals model colored by charge wtih FirstGlance in Jmol. Sidechain nitrogens on Arg/Lys; oxygens on Asp/Glu. |
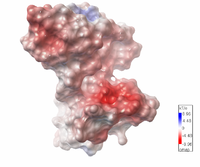 | Electrostatic potential map of 1tsj made with the Embedded Python Molecular Viewer from the Center for Computational Structural Biology of the Scripps Research Institute.
Click on the image to enlarge. |
See Also
- Electrostatic interactions in Proteopedia.
- Jmol/Electrostatic potential methods.
- Isopotential Map in Wikipedia
- Delphi Web Server
