We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 1845
From Proteopedia
(Difference between revisions)
| Line 23: | Line 23: | ||
=== Catalytic Triad === | === Catalytic Triad === | ||
[[Image:Final rayed image of binding pocket.png|400 px|right|thumb|Figure 1: Ser, His, Asp catalytic triad non-covalent stabilizing interactions with oxyanion hole.]] | [[Image:Final rayed image of binding pocket.png|400 px|right|thumb|Figure 1: Ser, His, Asp catalytic triad non-covalent stabilizing interactions with oxyanion hole.]] | ||
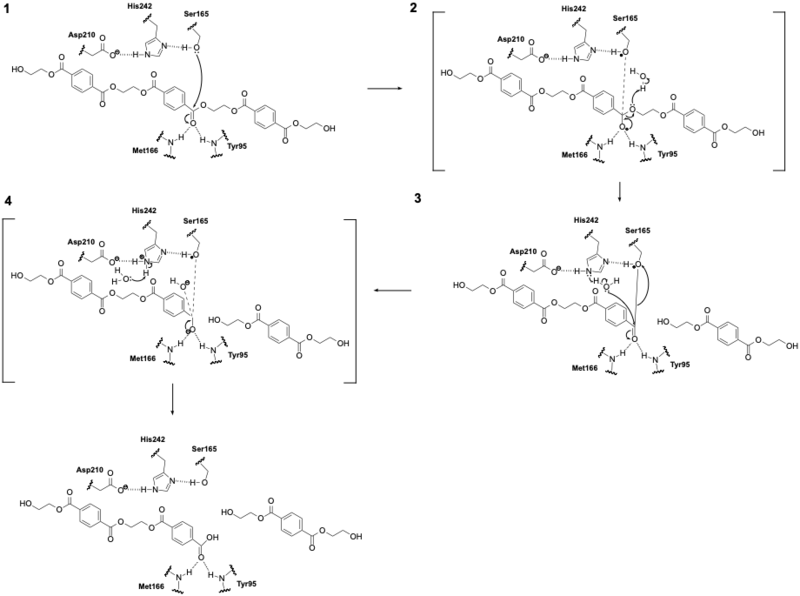
| - | LCC catalyzes the breakdown of PET using a serine hydrolase mechanism with a <scene name='10/1075247/Catalytic_triad3_w_label/3'>catalytic triad</scene> of Ser165, | + | LCC catalyzes the breakdown of PET using a serine hydrolase mechanism with a <scene name='10/1075247/Catalytic_triad3_w_label/3'>catalytic triad</scene> of Ser165, Asp210, and His242. (Figure 1) (1) The reaction begins when His242 deprotonates Ser165, which activates it as a nucleophile. (2) Ser165 then attacks the carbonyl carbon of an ester bond in the PET polymer to form a tetrahedral intermediate. (3) This tetrahedral intermediate is stabilized by an oxyanion hole formed by the backbone amides of Met166 and Tyr95. (4) The intermediate collapses; one product is released and an acyl-enzyme intermediate is formed. (5) A water molecule, activated by His242, then attacks the acyl-enzyme. This releases the second product and resets the enzyme’s active site. |
[[Image:Mech2.png|800 px|right|thumb|Figure 2: LCC mechanism. LCC hydrolyzes PET using a catalytic triad (Ser165, His242, Asp210) to cleave ester bonds via a tetrahedral intermediate.]] | [[Image:Mech2.png|800 px|right|thumb|Figure 2: LCC mechanism. LCC hydrolyzes PET using a catalytic triad (Ser165, His242, Asp210) to cleave ester bonds via a tetrahedral intermediate.]] | ||
| Line 36: | Line 36: | ||
From this screen, two mutations at Phe243 (F243I and F243W) were shown to improve catalytic activity by optimizing substrate positioning within the groove. To increase thermostability, the authors targeted a region of the enzyme that is structurally analogous to known divalent [https://en.wikipedia.org/wiki/Metal-binding_protein metal binding] sites in other cutinases. Instead of using stabilizing ions, which could complicate industrial degradation processes, they engineered a [https://en.wikipedia.org/wiki/Disulfide disulfide bridge] by mutating Asp238 and Ser283 to Cys residues (D238C/S283C). Additional mutations were selected based on thermostability screening. Among these, Y127G improved the melting point without reducing activity. | From this screen, two mutations at Phe243 (F243I and F243W) were shown to improve catalytic activity by optimizing substrate positioning within the groove. To increase thermostability, the authors targeted a region of the enzyme that is structurally analogous to known divalent [https://en.wikipedia.org/wiki/Metal-binding_protein metal binding] sites in other cutinases. Instead of using stabilizing ions, which could complicate industrial degradation processes, they engineered a [https://en.wikipedia.org/wiki/Disulfide disulfide bridge] by mutating Asp238 and Ser283 to Cys residues (D238C/S283C). Additional mutations were selected based on thermostability screening. Among these, Y127G improved the melting point without reducing activity. | ||
| - | These mutations were combined to create multi-mutant LCC variants with improved activity and thermostability. The two most successful variants were: | ||
| - | ICCG: F243I / D238C / S283C / Y127G | ||
| - | + | === F243 === | |
| - | + | <scene name='10/1075247/F243_original/1'>F243</scene> is located 3.6 Å from the ligand. Two mutations at this position, <scene name='10/1075247/F243i/1'>F243I</scene> and F243W, increase the catalytic activity of the enzyme. The F243I mutation inserts the smaller isoleucine whose side chain allows the ligand to sit closer. This reduces the ligand distance to 3.0 Å, improving substrate binding. The F243W mutation inserts the bulkier, nitrogen-containing aromatic aide chain. Trp brings the ligand slightly closer at 3.2 Å and introduces potential for new interactions, such as hydrogen bonding or [https://en.wikipedia.org/wiki/Pi-stacking#:~:text=In%20chemistry%2C%20pi%20stacking%20(also,interaction%22)%20is%20electrostatically%20repulsive. π-stacking]. Both mutations result in improved catalytic performance. The F243I mutant shows a 27.5% increase in activity, while the F243W mutant shows a 17.5% increase, compared to the wild-type enzyme.<ref name="Tournier"/> | |
| - | + | ||
| - | + | ||
| - | + | ||
=== Tyr 127 === | === Tyr 127 === | ||
| - | The mutation of Tyr to Gly at <scene name='10/1075247/Y127/4'> | + | The mutation of Tyr to Gly at <scene name='10/1075247/Y127/4'>Tyr127</scene> also increases the thermostability. The melting point of Y127G is increased to 87.0°C from the WT melting point of 84.7°C. Tyr has a bulky, rigid aromatic side chain that can cause structural strain, shown in <scene name='10/1075247/Y127_spacefill/1'>Y127 representation</scene>. Gly is the smallest amino acid and lacks a side chain, providing greater flexibility. The mutation <scene name='10/1075247/Y127g/2'>Y127G</scene> melting point is increased to 87.0°C. The mutation reduces [https://en.wikipedia.org/wiki/Steric_effects#:~:text=Steric%20hindrance%20is%20the%20slowing,as%20slowing%20unwanted%20side%2Dreactions. steric hindrance] and relieves strain in the protein structure, demonstrated in <scene name='10/1075247/Y127g_space_fill/1'>Y127G representation</scene>. By increasing flexibility, the Y127G mutation helps the protein maintain its folded structure under heat stress.<ref name="Tournier"/> |
=== Ser 283 & Asp 238 === | === Ser 283 & Asp 238 === | ||
Revision as of 19:55, 22 April 2025
| This Sandbox is Reserved from March 18 through September 1, 2025 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson and Mark Macbeth at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1828 through Sandbox Reserved 1846. |
To get started:
More help: Help:Editing |
Leaf Branch Compost Cutinase
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 Tournier V, Topham CM, Gilles A, David B, Folgoas C, Moya-Leclair E, Kamionka E, Desrousseaux ML, Texier H, Gavalda S, Cot M, Guemard E, Dalibey M, Nomme J, Cioci G, Barbe S, Chateau M, Andre I, Duquesne S, Marty A. An engineered PET depolymerase to break down and recycle plastic bottles. Nature. 2020 Apr;580(7802):216-219. doi: 10.1038/s41586-020-2149-4. Epub 2020 Apr, 8. PMID:32269349 doi:http://dx.doi.org/10.1038/s41586-020-2149-4
- ↑ 2.0 2.1 2.2 2.3 2.4 Sui B, Wang T, Fang J, Hou Z, Shu T, Lu Z, Liu F, Zhu Y. Recent advances in the biodegradation of polyethylene terephthalate with cutinase-like enzymes. Front Microbiol. 2023 Oct 2;14:1265139. PMID:37849919 doi:10.3389/fmicb.2023.1265139
- ↑ Ueda H, Tabata J, Seshime Y, Masaki K, Sameshima-Yamashita Y, Kitamoto H. Cutinase-like biodegradable plastic-degrading enzymes from phylloplane yeasts have cutinase activity. Biosci Biotechnol Biochem. 2021 Jul 23;85(8):1890-1898. PMID:34160605 doi:10.1093/bbb/zbab113
- ↑ Kolattukudy PE. Biopolyester membranes of plants: cutin and suberin. Science. 1980 May 30;208(4447):990-1000. PMID:17779010 doi:10.1126/science.208.4447.990
- ↑ 5.0 5.1 5.2 5.3 5.4 Khairul Anuar NFS, Huyop F, Ur-Rehman G, Abdullah F, Normi YM, Sabullah MK, Abdul Wahab R. An Overview into Polyethylene Terephthalate (PET) Hydrolases and Efforts in Tailoring Enzymes for Improved Plastic Degradation. Int J Mol Sci. 2022 Oct 20;23(20):12644. PMID:36293501 doi:10.3390/ijms232012644
- ↑ 6.0 6.1 Burgin T, Pollard BC, Knott BC, Mayes HB, Crowley MF, McGeehan JE, Beckham GT, Woodcock HL. The reaction mechanism of the Ideonella sakaiensis PETase enzyme. Commun Chem. 2024 Mar 27;7(1):65. PMID:38538850 doi:10.1038/s42004-024-01154-x
- ↑ 7.0 7.1 7.2 Zhang J, Wang H, Luo Z, Yang Z, Zhang Z, Wang P, Li M, Zhang Y, Feng Y, Lu D, Zhu Y. Computational design of highly efficient thermostable MHET hydrolases and dual enzyme system for PET recycling. Commun Biol. 2023 Nov 9;6(1):1135. PMID:37945666 doi:10.1038/s42003-023-05523-5
- ↑ Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K. A bacterium that degrades and assimilates poly(ethylene terephthalate). Science. 2016 Mar 11;351(6278):1196-9. doi: 10.1126/science.aad6359. PMID:26965627 doi:http://dx.doi.org/10.1126/science.aad6359
- ↑ Landrigan PJ, Stegeman JJ, Fleming LE, Allemand D, Anderson DM, Backer LC, Brucker-Davis F, Chevalier N, Corra L, Czerucka D, Bottein MD, Demeneix B, Depledge M, Deheyn DD, Dorman CJ, Fénichel P, Fisher S, Gaill F, Galgani F, Gaze WH, Giuliano L, Grandjean P, Hahn ME, Hamdoun A, Hess P, Judson B, Laborde A, McGlade J, Mu J, Mustapha A, Neira M, Noble RT, Pedrotti ML, Reddy C, Rocklöv J, Scharler UM, Shanmugam H, Taghian G, van de Water JAJM, Vezzulli L, Weihe P, Zeka A, Raps H, Rampal P. Human Health and Ocean Pollution. Ann Glob Health. 2020 Dec 3;86(1):151. PMID:33354517 doi:10.5334/aogh.2831
- ↑ Jambeck JR, Geyer R, Wilcox C, Siegler TR, Perryman M, Andrady A, Narayan R, Law KL. Marine pollution. Plastic waste inputs from land into the ocean. Science. 2015 Feb 13;347(6223):768-71. PMID:25678662 doi:10.1126/science.1260352
Student Contributors
Ashley Callaghan, Rebecca Hoff, & Simone McCowan