We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
User:Marcos Ngo/Sandbox 1
From Proteopedia
(Difference between revisions)
| Line 4: | Line 4: | ||
DNA glycosylases search the entire genome for DNA lesions. These highly selective enzymes recognize a damaged base and remove it. There are four super families: Udg, Nth, Nei, and AGG. DNA glycosylases are used to bind to and excise the base. First, the DNA is “pinched” by the enzyme, which destabilizes the helix. From here, they use a wedge amino acid to “push” the lesion out of the helix. While the lesion is being flipped out, another amino acid “plugs” into the helix to fill the gap and maintain the structure of the helix. Finally, the lesion is “pulled” into the active site to allow for lesion removal. This has been termed the “pinch, push, plug, and pull” mechanism for base flipping.<ref>https://scholarworks.uvm.edu/cgi/viewcontent.cgi?article=2160&context=graddis</ref><ref>PMID:20469926</ref><ref>PMID:12220189</ref><ref>PMID:12220189</ref> | DNA glycosylases search the entire genome for DNA lesions. These highly selective enzymes recognize a damaged base and remove it. There are four super families: Udg, Nth, Nei, and AGG. DNA glycosylases are used to bind to and excise the base. First, the DNA is “pinched” by the enzyme, which destabilizes the helix. From here, they use a wedge amino acid to “push” the lesion out of the helix. While the lesion is being flipped out, another amino acid “plugs” into the helix to fill the gap and maintain the structure of the helix. Finally, the lesion is “pulled” into the active site to allow for lesion removal. This has been termed the “pinch, push, plug, and pull” mechanism for base flipping.<ref>https://scholarworks.uvm.edu/cgi/viewcontent.cgi?article=2160&context=graddis</ref><ref>PMID:20469926</ref><ref>PMID:12220189</ref><ref>PMID:12220189</ref> | ||
| - | hNTHL1 or human Endonuclease III (Nth) is a 36 kDa bifunctional DNA glycosylase involved in the base excision repair process. A bifunctional glycosylase refers to the ability of the protein to be able to recognize and '''excise''' damaged bases from DNA and cleave the DNA backbone at the abasic site. This enzyme has a preference for oxidized pyrimidines with Tg (Thymine Glycol) being the preferred substrate. Upon encountering this damaged base, the protein cleaves the N-glycosidic bond, which leaves an apurinic site. From here, the backbone is cleaved via beta elimination, which leaves a 3’ aldehyde and creates a single-strand break. Next, the DNA is handed off to Apurinic endonuclease 1 or polynucleotide kinase, leaving a free 3′ hydroxyl for DNA polymerase β to insert the correct nucleotide. Finally, the nick is sealed by the DNA ligase IIIα. <ref>PMID:34871433</ref><ref>PMID:20005182</ref><ref>PMID:9295348</ref> | + | hNTHL1 or human Endonuclease III (Nth) is a 36 kDa bifunctional DNA glycosylase involved in the base excision repair process. A bifunctional glycosylase refers to the ability of the protein to be able to recognize and '''excise''' damaged bases from DNA and cleave the DNA backbone at the abasic site. This enzyme has a preference for oxidized pyrimidines with, Tg (Thymine Glycol) being the preferred substrate. Upon encountering this damaged base, the protein cleaves the N-glycosidic bond, which leaves an apurinic site. From here, the backbone is cleaved via beta elimination, which leaves a 3’ aldehyde and creates a single-strand break. Next, the DNA is handed off to Apurinic endonuclease 1 or polynucleotide kinase, leaving a free 3′ hydroxyl for DNA polymerase β to insert the correct nucleotide. Finally, the nick is sealed by the DNA ligase IIIα. <ref>PMID:34871433</ref><ref>PMID:20005182</ref><ref>PMID:9295348</ref> |
== Structural Highlights == | == Structural Highlights == | ||
| - | This enzyme hNTHL1 belongs to the HhH (Helix-Hairpin-Helix) superfamily which consists of <scene name='10/1077482/Two_domains/2'>two alpha-helical domains</scene> which are connected by two linkers. The solved structure of hNTHl1, with the first 63 residues being removed due to disorder, reveals <scene name='10/1077482/Ncfedomain1/3'>Domain 1</scene> with the iron sulfur cluster, N- and C-termini, and a catalytic resiude (asp). <scene name='10/1077482/Domain2features/2'>Domain 2</scene> has six helical barrels, hairpin-helix-hairpin, and the final catalytic residue (gly). The <scene name='10/1077482/Proglyhhh/1'>HhH</scene> motif has characteristic glycine and proline rich loop. The HhH allows for hydrogen bond interactions with the DNA backbone. <ref>PMID:12840008</ref><ref>https://scholarworks.uvm.edu/cgi/viewcontent.cgi?article=2160&context=graddis</ref><ref>PMID:1283262</ref> | + | This enzyme hNTHL1 belongs to the HhH (Helix-Hairpin-Helix) superfamily which consists of <scene name='10/1077482/Two_domains/2'>two alpha-helical domains</scene>, which are connected by two linkers. The solved structure of hNTHl1, with the first 63 residues being removed due to disorder, reveals <scene name='10/1077482/Ncfedomain1/3'>Domain 1</scene> with the iron sulfur cluster, N- and C-termini, and a catalytic resiude (asp). <scene name='10/1077482/Domain2features/2'>Domain 2</scene> has six helical barrels, hairpin-helix-hairpin, and the final catalytic residue (gly). The <scene name='10/1077482/Proglyhhh/1'>HhH</scene> motif has a characteristic glycine and proline-rich loop. The HhH allows for hydrogen bond interactions with the DNA backbone. <ref>PMID:12840008</ref><ref>https://scholarworks.uvm.edu/cgi/viewcontent.cgi?article=2160&context=graddis</ref><ref>PMID:1283262</ref> |
This structure is captured in an <scene name='10/1077482/Open_conformation/1'>open conformation</scene> where the catalytic residues Lys220 and Asp239 are positioned approximately 25 Å apart, which is too far for catalysis. This implies that a conformational change is required to assemble the active site. To find the closed conformation, an engineered chimera was made by swapping the <scene name='10/1077482/Linker1/1'>flexible interdomain linker</scene> in human NTHL1 with a shorter, more rigid linker from a bacterial homolog. The <scene name='10/1077482/Chimera/1'>hNTHL1Δ63 chimera</scene> structure adopts a closed conformation where Lys220 and Asp239 are approximately 5 Å apart, which mimics the configuration seen in catalytically active homologs. <ref>PMID:34871433</ref> | This structure is captured in an <scene name='10/1077482/Open_conformation/1'>open conformation</scene> where the catalytic residues Lys220 and Asp239 are positioned approximately 25 Å apart, which is too far for catalysis. This implies that a conformational change is required to assemble the active site. To find the closed conformation, an engineered chimera was made by swapping the <scene name='10/1077482/Linker1/1'>flexible interdomain linker</scene> in human NTHL1 with a shorter, more rigid linker from a bacterial homolog. The <scene name='10/1077482/Chimera/1'>hNTHL1Δ63 chimera</scene> structure adopts a closed conformation where Lys220 and Asp239 are approximately 5 Å apart, which mimics the configuration seen in catalytically active homologs. <ref>PMID:34871433</ref> | ||
| Line 16: | Line 16: | ||
</table> | </table> | ||
| - | The role of the <scene name='10/1077482/Fes_proper/2'>FeS Cluster</scene> is highly debated. One of the views is that the cluster is involved in scanning for lesions. Researchers found that oxidizing the FeS cluster in hNTHL1 from [4Fe-4S]^2+ to [4Fe-4S]^3+ increases its binding to DNA. When a mismatch such as C:A is introduced, this can disrupt DNA charge transport not allowing electrons to travel along the helix. This could stop the reduction of [4Fe-4S]^3+ to [4Fe-4S]^2+ leaving Nth bound until all | + | The role of the <scene name='10/1077482/Fes_proper/2'>FeS Cluster</scene> is highly debated. One of the views is that the cluster is involved in scanning for lesions. Researchers found that oxidizing the FeS cluster in hNTHL1 from [4Fe-4S]^2+ to [4Fe-4S]^3+ increases its binding to DNA. When a mismatch such as C:A is introduced, this can disrupt DNA charge transport not allowing electrons to travel along the helix. This could stop the reduction of [4Fe-4S]^3+ to [4Fe-4S]^2+ leaving Nth bound until all lesions are removed. Another view is that the FeS cluster plays a role as a structural scaffold to stabilize the interaction of the protein with the DNA. <ref>PMID:19720997</ref><ref>PMID:28817778</ref><ref>DOI:https://pubs.rsc.org/en/content/articlelanding/2022/cc/d2cc03643f</ref>. |
| - | The function of the N-terminus of hNHTL1 has been a subject of study. It is theorized that the N-terminus, which is extended compared to homologs, functions as a means to remain bound to DNA, protecting the labile abasic site while waiting for its handoff with APE1. This | + | The function of the N-terminus of hNHTL1 has been a subject of study. It is theorized that the N-terminus, which is extended compared to homologs, functions as a means to remain bound to DNA, protecting the labile abasic site while waiting for its handoff with APE1. This was found through truncation of the N-terminal region (residues 1-96), which revealed that deletion of 55, 75, or 80 residues from the N-terminus resulted in a four to fivefold increase in catalytic activity. The rate-limiting step in hNTHL1's reaction is the release of the free 3’ aldehyde. <ref>PMID:12144783</ref> |
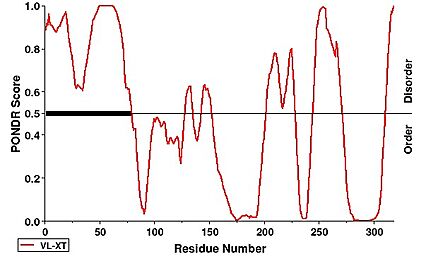
Notability, the first 63 residues were not modeled within the structure of hNHTL1 which is due to disorder. This can be observed under a PONDR prediction. | Notability, the first 63 residues were not modeled within the structure of hNHTL1 which is due to disorder. This can be observed under a PONDR prediction. | ||
Revision as of 18:27, 25 April 2025
Human NTHL1
| |||||||||||