This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2clt
From Proteopedia
| Line 1: | Line 1: | ||
[[Image:2clt.gif|left|200px]] | [[Image:2clt.gif|left|200px]] | ||
| - | + | <!-- | |
| - | + | The line below this paragraph, containing "STRUCTURE_2clt", creates the "Structure Box" on the page. | |
| - | + | You may change the PDB parameter (which sets the PDB file loaded into the applet) | |
| - | + | or the SCENE parameter (which sets the initial scene displayed when the page is loaded), | |
| - | + | or leave the SCENE parameter empty for the default display. | |
| - | | | + | --> |
| - | | | + | {{STRUCTURE_2clt| PDB=2clt | SCENE= }} |
| - | + | ||
| - | + | ||
| - | }} | + | |
'''CRYSTAL STRUCTURE OF THE ACTIVE FORM (FULL-LENGTH) OF HUMAN FIBROBLAST COLLAGENASE.''' | '''CRYSTAL STRUCTURE OF THE ACTIVE FORM (FULL-LENGTH) OF HUMAN FIBROBLAST COLLAGENASE.''' | ||
| Line 30: | Line 27: | ||
[[Category: Nagase, H.]] | [[Category: Nagase, H.]] | ||
[[Category: Visse, R.]] | [[Category: Visse, R.]] | ||
| - | [[Category: | + | [[Category: Autocatalytic cleavage]] |
| - | [[Category: | + | [[Category: Calcium]] |
| - | [[Category: | + | [[Category: Collagen]] |
| - | [[Category: | + | [[Category: Collagen degradation]] |
| - | [[Category: | + | [[Category: Extracellular matrix]] |
| - | [[Category: | + | [[Category: Fibroblast collagenase]] |
| - | [[Category: | + | [[Category: Glycoprotein]] |
| - | [[Category: | + | [[Category: Hydrolase]] |
| - | [[Category: | + | [[Category: Inhibitor-free]] |
| - | [[Category: | + | [[Category: Matrix metalloproteinase]] |
| - | [[Category: | + | [[Category: Metal-binding]] |
| - | [[Category: | + | [[Category: Metalloprotease]] |
| - | [[Category: | + | [[Category: Polymorphism]] |
| - | [[Category: | + | [[Category: Protease]] |
| - | [[Category: | + | [[Category: Zinc]] |
| - | [[Category: | + | [[Category: Zymogen]] |
| - | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Sat May 3 22:27:11 2008'' | |
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | + | |
Revision as of 19:27, 3 May 2008
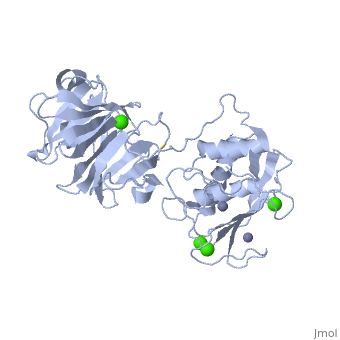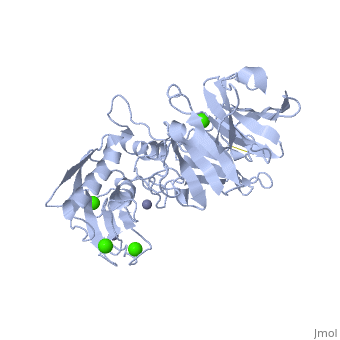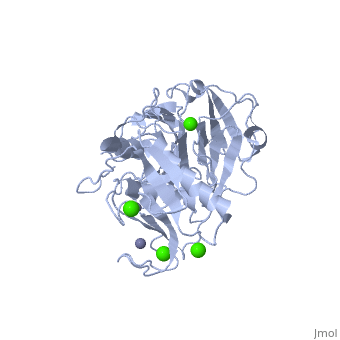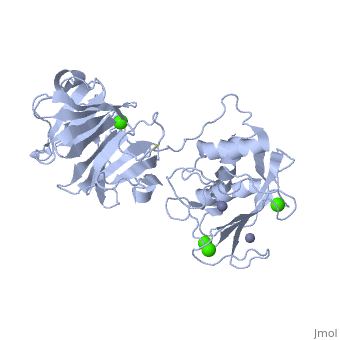
CRYSTAL STRUCTURE OF THE ACTIVE FORM (FULL-LENGTH) OF HUMAN FIBROBLAST COLLAGENASE.
Overview
The extracellular matrix is a dynamic environment that constantly undergoes remodelling and degradation during vital physiological processes such as angiogenesis, wound healing, and development. Unbalanced extracellular matrix breakdown is associated with many diseases such as arthritis, cancer and fibrosis. Interstitial collagen is degraded by matrix metalloproteinases with collagenolytic activity by MMP-1, MMP-8 and MMP-13, collectively known as the collagenases. Matrix metalloproteinase 1 (MMP-1) plays a pivotal role in degradation of interstitial collagen types I, II, and III. Here, we report the crystal structure of the active form of human MMP-1 at 2.67 A resolution. This is the first MMP-1 structure that is free of inhibitor and a water molecule essential for peptide hydrolysis is observed coordinated with the active site zinc. Comparing this structure with the human proMMP-1 shows significant structural differences, mainly in the relative orientation of the hemopexin domain, between the pro form and active form of the human enzyme.
About this Structure
2CLT is a Single protein structure of sequence from Homo sapiens. Full crystallographic information is available from OCA.
Reference
Crystal structure of an active form of human MMP-1., Iyer S, Visse R, Nagase H, Acharya KR, J Mol Biol. 2006 Sep 8;362(1):78-88. Epub 2006 Aug 4. PMID:16890240 Page seeded by OCA on Sat May 3 22:27:11 2008
Categories: Homo sapiens | Interstitial collagenase | Single protein | Acharya, K R. | Iyer, S. | Nagase, H. | Visse, R. | Autocatalytic cleavage | Calcium | Collagen | Collagen degradation | Extracellular matrix | Fibroblast collagenase | Glycoprotein | Hydrolase | Inhibitor-free | Matrix metalloproteinase | Metal-binding | Metalloprotease | Polymorphism | Protease | Zinc | Zymogen