This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1a0a
From Proteopedia
| Line 1: | Line 1: | ||
| - | [[Image:1a0a. | + | {{Seed}} |
| + | [[Image:1a0a.png|left|200px]] | ||
<!-- | <!-- | ||
| Line 9: | Line 10: | ||
{{STRUCTURE_1a0a| PDB=1a0a | SCENE= }} | {{STRUCTURE_1a0a| PDB=1a0a | SCENE= }} | ||
| - | + | ===PHOSPHATE SYSTEM POSITIVE REGULATORY PROTEIN PHO4/DNA COMPLEX=== | |
| - | + | <!-- | |
| - | The | + | The line below this paragraph, {{ABSTRACT_PUBMED_9303313}}, adds the Publication Abstract to the page |
| + | (as it appears on PubMed at http://www.pubmed.gov), where 9303313 is the PubMed ID number. | ||
| + | --> | ||
| + | {{ABSTRACT_PUBMED_9303313}} | ||
==About this Structure== | ==About this Structure== | ||
| Line 32: | Line 36: | ||
[[Category: Basic helix loop helix]] | [[Category: Basic helix loop helix]] | ||
[[Category: Transcription factor]] | [[Category: Transcription factor]] | ||
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on | + | |
| + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Mon Jun 30 15:42:46 2008'' | ||
Revision as of 12:42, 30 June 2008
| |||||||||
| 1a0a, resolution 2.80Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
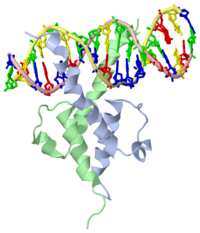
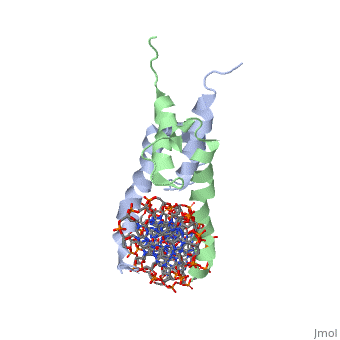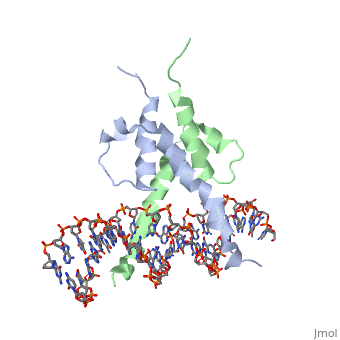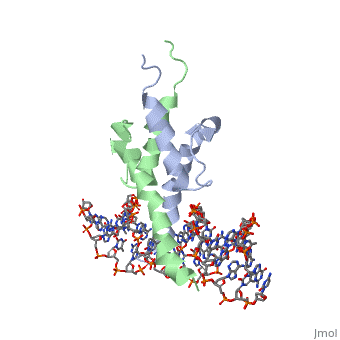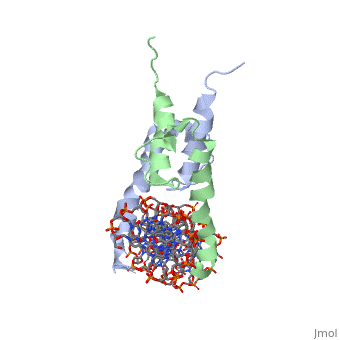
PHOSPHATE SYSTEM POSITIVE REGULATORY PROTEIN PHO4/DNA COMPLEX
The crystal structure of a DNA-binding domain of PHO4 complexed with DNA at 2.8 A resolution revealed that the domain folds into a basic-helix-loop-helix (bHLH) motif with a long but compact loop that contains a short alpha-helical segment. This helical structure positions a tryptophan residue into an aromatic cluster so as to make the loop compact. PHO4 binds to DNA as a homodimer with direct reading of both the core E-box sequence CACGTG and its 3'-flanking bases. The 3'-flanking bases GG are recognized by Arg2 and His5. The residues involved in the E-box recognition are His5, Glu9 and Arg13, as already reported for bHLH/Zip proteins MAX and USF, and are different from those recognized by bHLH proteins MyoD and E47, although PHO4 is a bHLH protein.
Crystal structure of PHO4 bHLH domain-DNA complex: flanking base recognition., Shimizu T, Toumoto A, Ihara K, Shimizu M, Kyogoku Y, Ogawa N, Oshima Y, Hakoshima T, EMBO J. 1997 Aug 1;16(15):4689-97. PMID:9303313
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
About this Structure
1A0A is a Single protein structure of sequence from Saccharomyces cerevisiae. Full crystallographic information is available from OCA.
Reference
Crystal structure of PHO4 bHLH domain-DNA complex: flanking base recognition., Shimizu T, Toumoto A, Ihara K, Shimizu M, Kyogoku Y, Ogawa N, Oshima Y, Hakoshima T, EMBO J. 1997 Aug 1;16(15):4689-97. PMID:9303313
Page seeded by OCA on Mon Jun 30 15:42:46 2008