This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1sox
From Proteopedia
| |||||||
| , resolution 1.9Å | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | , , , , and | ||||||
| Activity: | Sulfite oxidase, with EC number 1.8.3.1 | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
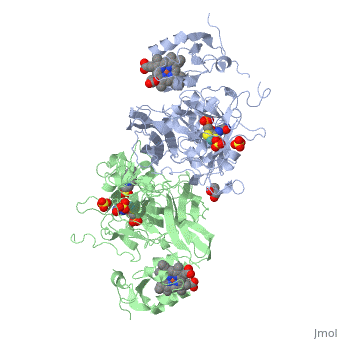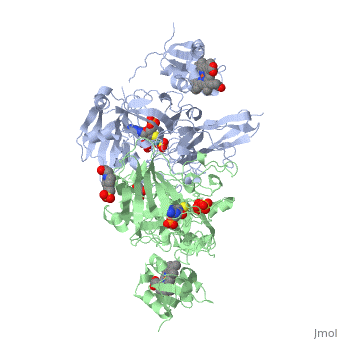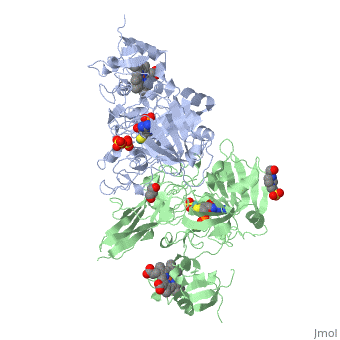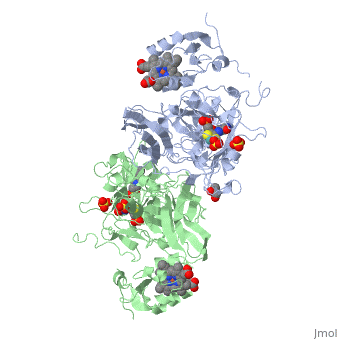
SULFITE OXIDASE FROM CHICKEN LIVER
Overview
The molybdenum-containing enzyme sulfite oxidase catalyzes the conversion of sulfite to sulfate, the terminal step in the oxidative degradation of cysteine and methionine. Deficiency of this enzyme in humans usually leads to major neurological abnormalities and early death. The crystal structure of chicken liver sulfite oxidase at 1.9 A resolution reveals that each monomer of the dimeric enzyme consists of three domains. At the active site, the Mo is penta-coordinated by three sulfur ligands, one oxo group, and one water/hydroxo. A sulfate molecule adjacent to the Mo identifies the substrate binding pocket. Four variants associated with sulfite oxidase deficiency have been identified: two mutations are near the sulfate binding site, while the other mutations occur within the domain mediating dimerization.
About this Structure
1SOX is a Single protein structure of sequence from Gallus gallus. Full crystallographic information is available from OCA.
Reference
Molecular basis of sulfite oxidase deficiency from the structure of sulfite oxidase., Kisker C, Schindelin H, Pacheco A, Wehbi WA, Garrett RM, Rajagopalan KV, Enemark JH, Rees DC, Cell. 1997 Dec 26;91(7):973-83. PMID:9428520
Page seeded by OCA on Thu Mar 20 14:07:47 2008
Categories: Gallus gallus | Single protein | Sulfite oxidase | Kisker, C. | Rees, D C. | Schindelin, H. | EPE | GOL | HEM | MO | MTE | SO4 | Oxidoreductase | Sulfite oxidation