This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2adr
From Proteopedia
| |||||||
| Sites: | |||||||
| Ligands: | |||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
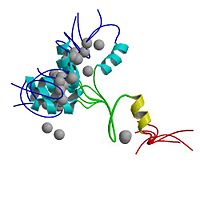
ADR1 DNA-BINDING DOMAIN FROM SACCHAROMYCES CEREVISIAE, NMR, 25 STRUCTURES
Overview
The region responsible for sequence-specific DNA binding by the transcription factor ADR1 contains two Cys2-His2 zinc fingers and an additional N-terminal proximal accessory region (PAR). The N-terminal (non-finger) PAR is unstructured in the absence of DNA and undergoes a folding transition on binding the DNA transcription target site. We have used a set of HN-HN NOEs derived from a perdeuterated protein-DNA complex to describe the fold of ADR1 bound to the UAS1 binding site. The PAR forms a compact domain consisting of three antiparallel strands that contact A-T base pairs in the major groove. The three-strand domain is a novel fold among all known DNA-binding proteins. The PAR shares sequence homology with the N-terminal regions of other zinc finger proteins, suggesting that it represents a new DNA-binding module that extends the binding repertoire of zinc finger proteins.
About this Structure
2ADR is a Single protein structure of sequence from Saccharomyces cerevisiae. Full crystallographic information is available from OCA.
Reference
A folding transition and novel zinc finger accessory domain in the transcription factor ADR1., Bowers PM, Schaufler LE, Klevit RE, Nat Struct Biol. 1999 May;6(5):478-85. PMID:10331877
Page seeded by OCA on Thu Mar 20 15:48:24 2008