This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1fxx
From Proteopedia
|
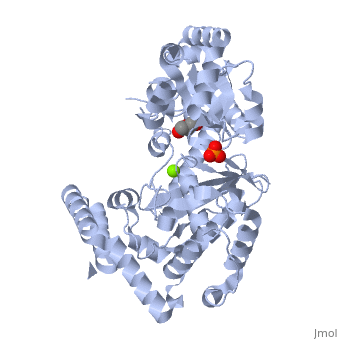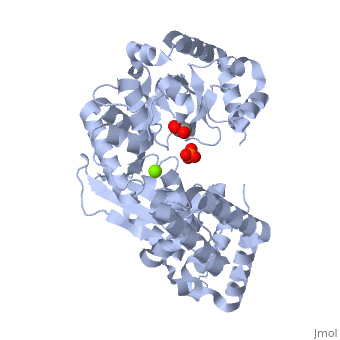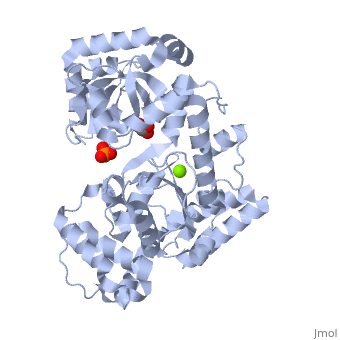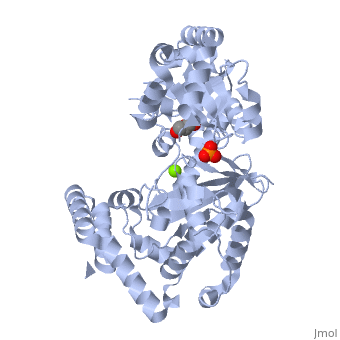
THE STRUCTURE OF EXONUCLEASE I SUGGESTS HOW PROCESSIVITY IS ACHIEVED
Overview
Exonuclease I (ExoI) from Escherichia coli is a monomeric enzyme that processively degrades single stranded DNA in the 3' to 5' direction and has been implicated in DNA recombination and repair. Determination of the structure of ExoI to 2.4 A resolution using X-ray crystallography verifies the expected correspondence between a region of ExoI and the exonuclease (or proofreading) domains of the DNA polymerases. The overall fold of ExoI also includes two other regions, one of which extends the exonuclease domain and another that can be described as an elaborated SH3 domain. These three regions combine to form a molecule that is shaped like the letter C, although it also contains a segment that effectively converts the C into an O-like shape. The structure of ExoI thus provides additional support for the idea that DNA metabolizing enzymes achieve processivity by completely enclosing the DNA.
About this Structure
1FXX is a Single protein structure of sequence from Escherichia coli with , and as ligands. Active as Exodeoxyribonuclease I, with EC number 3.1.11.1 Full crystallographic information is available from OCA.
Reference
Structure of Escherichia coli exonuclease I suggests how processivity is achieved., Breyer WA, Matthews BW, Nat Struct Biol. 2000 Dec;7(12):1125-8. PMID:11101894
Page seeded by OCA on Thu Feb 21 12:43:50 2008