This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox88
From Proteopedia
From Proteopedia Jump to: navigation, search
This sandbox is in use until August 1, 2011 for UMass Chemistry 423. Others please do not edit this page. Thanks!
Caspase 7
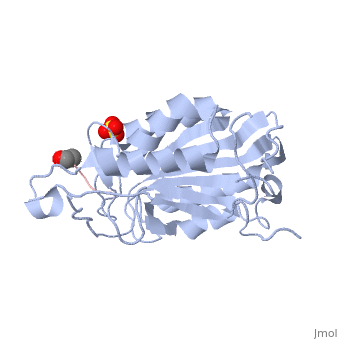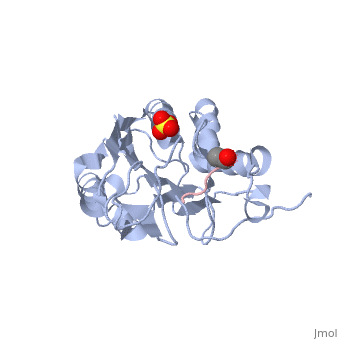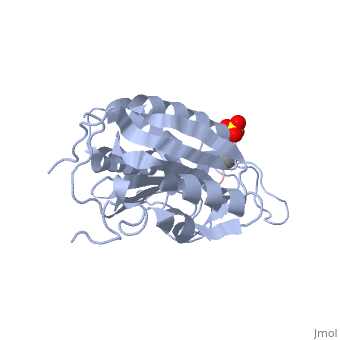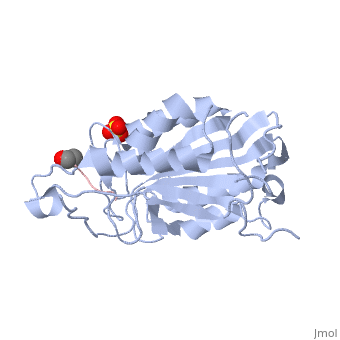
This is the structure for Caspase 7 with substrate bound. Caspase 7 is a Cysteine dependent protease that cleaves after ASPartate residues. The substrate here is the peptide DEVD. Caspase 7 plays a critical role in apoptosis as an initiator that cleaves downstream targets.
All caspases are zymogens. In the case of Casp7, this means that there is a prodomain that, when attached, renders the protein inactive. However, this prodomain is easily cleaved off, thus activating Casp7 as a protease.
| |||||||
| 1f1j, resolution 2.35Å () | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | |||||||
| Non-Standard Residues: | , | ||||||
| |||||||
| |||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
Active Site
The active site for Caspase 7 takes full advantage of acid base chemistry. Histidine 144 will act as a base, pulling the hydrogen away from cysteine 186. This will make that cysteine very nucleophilic and thus very active. It will bind substrate and cleave after an aspartate residue.