This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox20
From Proteopedia
Contents |
Introduction
Replication Termination in E. coli and B. subtilis
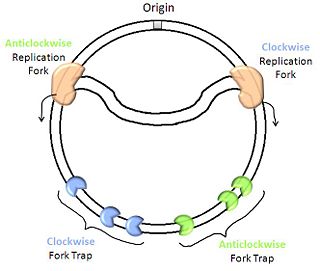
Replication forks proceed from the origin of the circular chromosome in opposite directions, creating a characteristic theta structure . Although the use of two active polymerase complexes is clearly faster than one, this mechanism generates difficulty when it comes to terminating replication. Among other factors, the presence of DNA-associated proteins causes the two replication complexes to proceed at different rates, and they therefore do not necessarily meet directly opposite the origin. However, the position of termination is not random, and is defined by the presence of termination sites which associate protein factors capable of halting the advance of the replication fork from only one direction. From their positions within the chromosome and their directionality, you can see that this occurs only once the complex has traversed more than half of the total DNA.
The Proteins
RTP
|
This is the area for text about RTP
Tus
|
This is the area for text about Tus
Tus binds to a conserved cytosine residue which is not base paired. The interactions between residues of the Tus protein and this unpaired cytosine nucleotide are .