This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1tyf
From Proteopedia
| |||||||||
| 1tyf, resolution 2.30Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Activity: | Endopeptidase Clp, with EC number 3.4.21.92 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
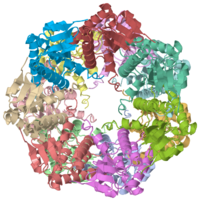
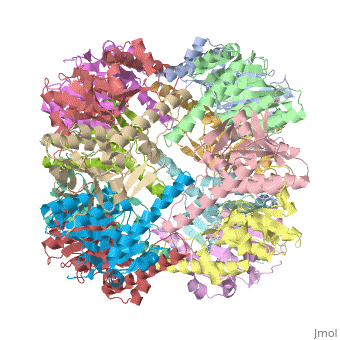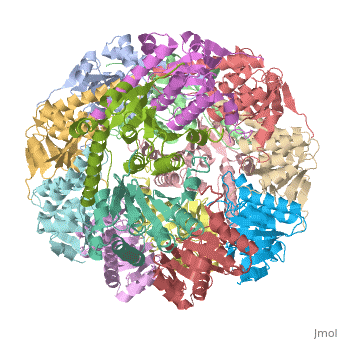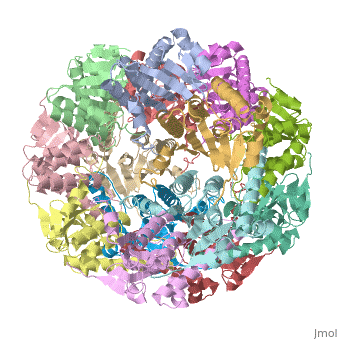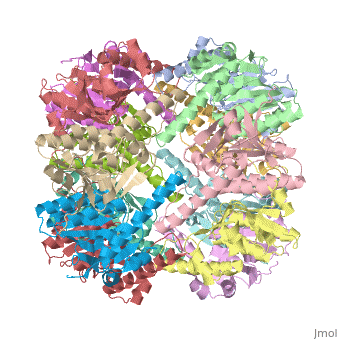
THE STRUCTURE OF CLPP AT 2.3 ANGSTROM RESOLUTION SUGGESTS A MODEL FOR ATP-DEPENDENT PROTEOLYSIS
We have determined the crystal structure of the proteolytic component of the caseinolytic Clp protease (ClpP) from E. coli at 2.3 A resolution using an ab initio phasing procedure that exploits the internal 14-fold symmetry of the oligomer. The structure of a ClpP monomer has a distinct fold that defines a fifth structural family of serine proteases but a conserved catalytic apparatus. The active protease resembles a hollow, solid-walled cylinder composed of two 7-fold symmetric rings stacked back-to-back. Its 14 proteolytic active sites are located within a central, roughly spherical chamber approximately 51 A in diameter. Access to the proteolytic chamber is controlled by two axial pores, each having a minimum diameter of approximately 10 A. From the structural features of ClpP, we suggest a model for its action in degrading proteins.
The structure of ClpP at 2.3 A resolution suggests a model for ATP-dependent proteolysis., Wang J, Hartling JA, Flanagan JM, Cell. 1997 Nov 14;91(4):447-56. PMID:9390554
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
About this Structure
1tyf is a 14 chain structure with sequence from Escherichia coli. Full crystallographic information is available from OCA.
See Also
Reference
- Wang J, Hartling JA, Flanagan JM. The structure of ClpP at 2.3 A resolution suggests a model for ATP-dependent proteolysis. Cell. 1997 Nov 14;91(4):447-56. PMID:9390554