The protein AmelASP1 has been identified in the antennae from the honeybee A.mellifera. Its primary sequence is a 144 amino acids polypeptide with a molecular weight of 13.180 kDa.
AmelASP1 is part of the Pheromone Binding Protein (PBP) family.
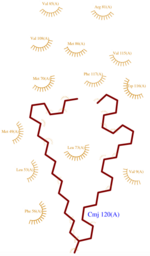
The 3D representation shown below was obtained at pH 5.5 using the nano-drops technique.
[1]
[2]
[3]
[4]
Updated on 23-December-2014
Biological function
Social relevance
Location in the antenna and transport of pheromones
Structure
Domains and family
The C terminal(scene) domain of this molecule presents a characteristic PBP-GOP domain. While this protein is composed of 144 residues the domain PBP begin at the 25th residue. AmelASP1 binds its ligand at low pH and releases it at neutral pH.
Key residues
AmelASP1 is composed of [5]
- : residues 8–25
- : residues 27–36
- : residues 42–56
- : residues 66–74
- : residues 75–77 (rarely mentionned in publications because of its tiny size)
- : residues 78–90
- : residues 96–112
has a break in the hydrogen-bonding pattern of its structure, forming tight substitute hydrogen bonds with water molecules. Thus, it results in a (at residue Ala 14) induced by a disruption in the helical conformation, due to hydrogen bonds with water molecules.
Components implicated in the structure rigidity
AmelASP1 presents which are greatly enhancing its structure’s rigidity by linking four of the helices together. The six cysteines and their interval spacing are the most striking features shared by proteins belonging to the PBP family.
The is established between and through Cysteins 20 and 51. links and through Cys 47 and 98, and the connects and thanks to Cys 89 and Cys 107.
Furthermore, non covalent bonds also play an important role.
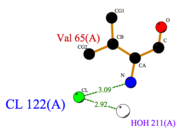
Indeed, at pH 5.5, among the numerous other, two hydrogene bonds are particularly noticeable because of their importance in the formation of the loop stabilizing H4. is established by Asp 66 and Leu 58 whereas is formed between Asp 60 and Ala 63.
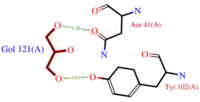
Cavity
The dynamic structure of the protein is responsible of the ligand’s binding by adjustment of position. The structure looses its flexibility when binds. The successful delivery of the effector to the receptor relies on this property. The ligand binding pocket consists in a cavity formed by the helices H2, H4 and H5(scene), arranged in a globular shape which leads to a clear separation of the
ligand from the hydrophilic environment.
The top of the cavity is not closed and can establish contacts with the solvent. The cavity is prone to accept ligand such as 9-ODA because of its specific composition. Indeed, cavity's components are mainly .They consequently interact with the ligand's hydrophobic carbon chain and are localized on the internal face of the helix.Thus, it implies that these residues respect a regular distance pattern in the primary structure of the AmelASP1.
pH influence
Ligands
Artificial ligands
Natural ligands
Related structures
These structures shown below representing the same protein with variable ligand and pH emphasize and illustrate the binding versatility of PBP.
- 3fe8 The same protein in complex with a serendipitous ligand soaked at pH 4.0
- 3fe9 The same protein in complex with a serendipitous ligand soaked at pH 7.0
- 3cdn The same protein in apo form soaked at pH 4.0
- 2h8v The same protein in apo form at pH 5.5
- 3cz2 The same protein in apo form at pH 7.0
- 3bfa The same protein in complex with the QMP at pH 5.5
- 3bfb The same protein in complex with the 9-ODA at pH 5.5
- 3bfh The same protein in complex with the HDOA at pH 5.5
- 3bjh The same protein in complex with the nBBSA at pH 5.5
- 3cyz The same protein in complex with the 9-ODA at pH 7.0
- 3cz0 The same protein in complex with the QMP at pH 7.0
- 3cz1 The same protein in complex with the nBBSA at pH 7.0
- 3cab The same protein in complex with the nBBSA soaked at pH 7.0
Relevance
Structural highlights
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.