This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Jennifer Taylor/Sandbox 6
From Proteopedia
Contents |
Overview
Throughout the course of this past year, we attempted to determine the function of our protein 2QRU. Protein shape and protein function are really closely related, so if two proteins that look similar are compared, there is a high likelihood that they do the same thing. This was the underlying basis for our project. After initial computer analysis, we found that many of the proteins that had structural similarity to 2QRU were esterases. Thus, our preliminary hypothesis predicted that 2QRU would be a lipase, a subclass of esterases. We performed an esterase assay to prove that 2QRU was an esterase first, and are yet to determine if it can be classified as a lipase.
Background
Proteins are one of four major macromolecules in biology. Present in nearly every living organism, proteins have a diverse set of functions ranging from regulating cell activity to catalyze reactions. Due to the sheer number of proteins in existence, there still remain many to discover and characterize. In 2000, the Protein Structure Initiative began an attempt to solve 3D-structures of proteins with known sequences in order to understand their functions. Though the initiative was successful, they faced financial drawbacks in 2015. There still remain several protein structures with unknown functions in a public database called the Protein Data Bank. In a final PSI summary published in 2017, 6920 structures had been solved in their seventeen years of work. What we tried to do is take one of the protein structures solved by the PSI and characterize its function.
Initial Process
The first thing we did was try to see if we could express our protein in a cell. We first tried to insert our plasmid containing 2QRU into DH5∂ cells but found that this cell didn't produce the T7 polymerase that was necessary to transcribe 2QRU. We then switched to E. Coli cells. We inserted our plasmid into E. Coli using a standard transformation protocol. We then measured the concentration of our plasmid and then ran an SDS page gel to determine If our plasmid was successfully expressed in our cells. After that, we purified our protein or essentially squeezed it out of our cells to get protein concentrate. We did this using His-Pur Nickel-NTA Spin columns. We put our cell extract into a column, added a specific buffer, and then centrifuged the column. All of the non-protein substances in the cell were filtered out to the bottom of the column in the first few washes. This meant that by our fifth or six wash, the substances coming through the column were our protein, 2QRU. We then ran a gel with our protein extract to see if it was successfully expressed. What we found, as expected, was that the band at our protein weight got stronger as the purification process went on, meaning the sample from our first wash had many bands signifying other proteins in the cell, not just ours. But, the sample from our last wash had 1 clean band at the expected weight of our protein, 33.8 kda.
Structural Analysis
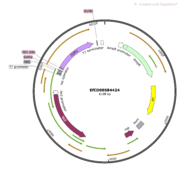
Figure 1 shows what my protein looks like inside a plasmid. We used SnapGene to figure out the weight of our protein 33.8 kda. Once we proved that we could produce our protein, we started to think about how we wanted to characterize its functions. Since we know that function and structure go hand in hand, we also used a program called PyMol to first look at the 3D structure of our protein. Here is a visualization of my protein highlighting the It’s an alpha/beta hydrolase that has one chain. It's 816 base pairs long. Then, using Dali and pFam, we ran a search for other proteins that were structurally similar to ours. We found a few hits including 1TAH and 1C4X that are classified as esterases by aligning the active sites of these proteins to ours.
| |||||||||||