This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1one
From Proteopedia
| |||||||||
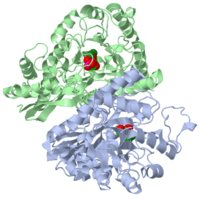
| 1one, resolution 1.80Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||||
| Non-Standard Residues: | |||||||||
| Activity: | Phosphopyruvate hydratase, with EC number 4.2.1.11 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
YEAST ENOLASE COMPLEXED WITH AN EQUILIBRIUM MIXTURE OF 2'-PHOSPHOGLYCEATE AND PHOSPHOENOLPYRUVATE
Template:ABSTRACT PUBMED 8605183
About this Structure
1ONE is a Single protein structure of sequence from Saccharomyces cerevisiae. Full crystallographic information is available from OCA.
Reference
A carboxylate oxygen of the substrate bridges the magnesium ions at the active site of enolase: structure of the yeast enzyme complexed with the equilibrium mixture of 2-phosphoglycerate and phosphoenolpyruvate at 1.8 A resolution., Larsen TM, Wedekind JE, Rayment I, Reed GH, Biochemistry. 1996 Apr 9;35(14):4349-58. PMID:8605183
Page seeded by OCA on Sun Jul 27 20:45:52 2008