This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Triose Phosphate Isomerase Structure & Mechanism
From Proteopedia
Triose Phosphate Isomerase (TIM)
This is a placeholder text to help you get started in placing a Jmol applet on your page. At any time, click "Show Preview" at the bottom of this page to see how it goes.
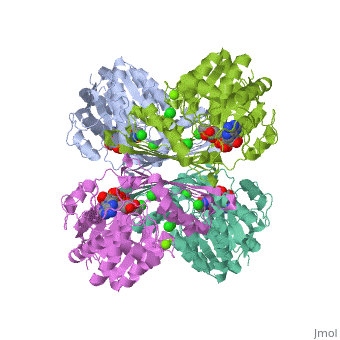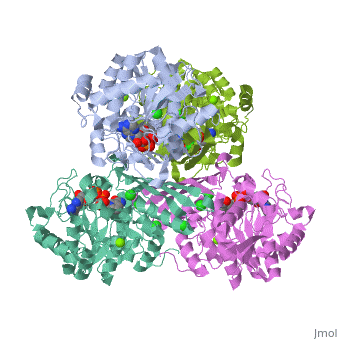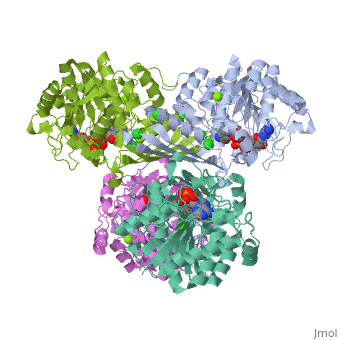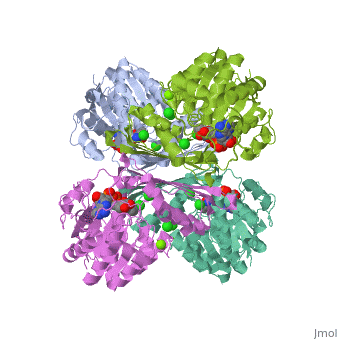
Triose phosphate isomerase (TIM) is a crucial enzyme in the glycolytic pathway. TIM reversibly converts the aldose Glyceraldehyde-3-phosphate (GASP) to the ketose Dihydroxyacetone phosphate (DHAP). The interconversion proceeds by an enediol intermediate.
Replace the PDB id (use lowercase!) after the STRUCTURE_ and after PDB= to load and display another structure.
| |||||||||
| 3cin, resolution 1.70Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , , | ||||||||
| Gene: | TM1419, TM_1419 (Thermotoga maritima MSB8) | ||||||||
| Activity: | Inositol-3-phosphate synthase, with EC number 5.5.1.4 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum, TOPSAN | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Proteopedia Page Contributors and Editors (what is this?)
Christian Krenk, Alexander Berchansky, Diamond B. Reese, Michal Harel, Jane S. Richardson, David Canner