This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 321
From Proteopedia
| This Sandbox is Reserved from January 10, 2010, through April 10, 2011 for use in BCMB 307-Proteins course taught by Andrea Gorrell at the University of Northern British Columbia, Prince George, BC, Canada. |
To get started:
More help: Help:Editing |
InhA
by Kelly Hrywkiw
| |||||||||
| 2h9i, resolution 2.20Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Gene: | inhA (Mycobacterium tuberculosis) | ||||||||
| Activity: | [acyl-carrier-protein_reductase_(NADH) Enoyl-[acyl-carrier-protein] reductase (NADH)], with EC number 1.3.1.9 | ||||||||
| Related: | 1zid | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
Introduction
The enzyme inhA is coded from the inhA gene that is simillar in sequence to the Salmonella typhimuriumgene which plays a role in fatty acid biosynthesis [1]. Inha is an NADH dependent trans enoyl-acyl ACP carrier protein that plays a role in the sysnthesis of Mycolic Acid, and is part of a short-chain dehydrogenase/reductase family [2][3]. Mycolic acids are long chain fatty acids that are essential in cell wall formation of the human pathogen Mycobacterium tuberculosisas well as other mycobateria such as Mycobacterium leprae[4]. Inha has been propsed as the target of the thionamide drugs, ethionamide (ETH) and isoniazid (INH), which have been used in treatment of mycobacterial infections [3].
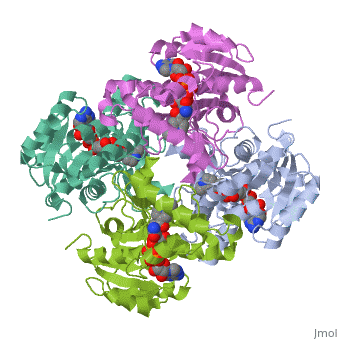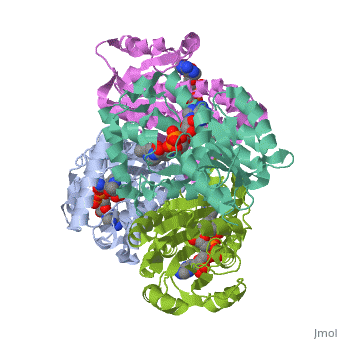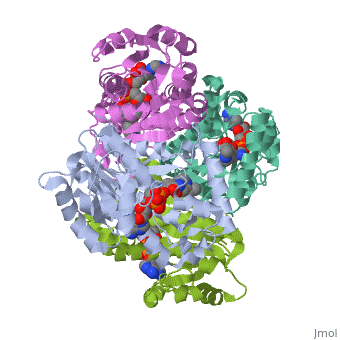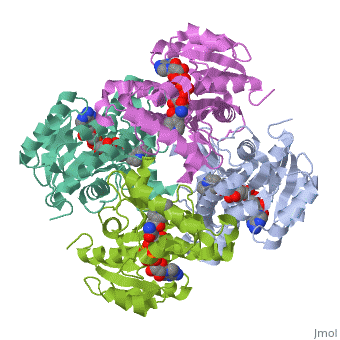
Structure of inhA
|
The overall strucuture of the inhA enzyme of Mycobacterium tuberculosis consists of a single domain with two substructures that are connected by short loop[1].
Substructure 1 of inhA
consists of 6 parallel β strands and 4 α helices interwoven together to form a core α/β structure that contains the n-terminal domain[1]. The first substructure can be further broken down into two sections, the consisting of two β strands and two short α helicies [1]. The first section is connected to the by a β strand that crosses over the two domains, and leads into the second section initiating at the third α helix [1]. (A-3) is connected by a long loop to a 14 residue β strand that then leads into the fourth α helix [1]. A-4 then leads into a fifth strand β , followed by a 25 residue α helix , and into the final strand β [1].
Substructure 2 of inhA
contains the c-terminal region of the molecule and consists of a small β strand (B-7), and two α helicies (A-6 and A-7) which are conected by a short five residue loop[1].
Physiological Function
Role in the Mycolic Acid Pathway
Protein Superfamilly
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 Strohmaier K, Streissle G, Clemm de Noronha S. [On the determination of size of early summer meningoencephalitis]. Arch Gesamte Virusforsch. 1965;17(2):300-3. PMID:5882878
- ↑ Wang F, Langley R, Gulten G, Dover LG, Besra GS, Jacobs WR Jr, Sacchettini JC. Mechanism of thioamide drug action against tuberculosis and leprosy. J Exp Med. 2007 Jan 22;204(1):73-8. Epub 2007 Jan 16. PMID:17227913 doi:10.1084/jem.20062100
- ↑ 3.0 3.1 Molle V, Gulten G, Vilcheze C, Veyron-Churlet R, Zanella-Cleon I, Sacchettini JC, Jacobs WR Jr, Kremer L. Phosphorylation of InhA inhibits mycolic acid biosynthesis and growth of Mycobacterium tuberculosis. Mol Microbiol. 2010 Dec;78(6):1591-605. doi:, 10.1111/j.1365-2958.2010.07446.x. Epub 2010 Nov 9. PMID:21143326 doi:10.1111/j.1365-2958.2010.07446.x
- ↑ . PMID:216315890657