TreX is an oligomer, as it exists in a dimeric state and a tetrameric state, both of which exhibit different enzymatic activities. All subunits are identical, where the monomer contains a total of 612 amino acids [1]. The polypeptide folds into two secondary structures, a β-sandwhich in the N terminal region, comprised of six β-strands and a (β/α)8 – barrel motif in the central domain, comprised of eight parallel α-strands which encircle eight parallel β-strands. The sequence composition of the TreX monomer exhibits a high degree of homology to the isoamylase debranching enzyme of Pseudomona, however the TreX monomer mainly deviates from this similarity in its substrate binding groove and the absence of a calcium ion ligand [1].
In the dimeric form, the individual subunits are adjacent to each other, where both of the active sites face the same side [1]. In the tetrameric form, two of the associated dimers face each other so as to position the substrate binding sites to the inside of the tetramer, facing each other with a slight offset [1]. Unlike other GDEs, TreX portrays different enzymatic activities in correlation to its oligomeric state. The dimeric form is predominantly associated with α-1,6-glycosidase, whereas the tetratmeic form is associated with α-1,4-transferase activity [1].
The active site of the TreX dimer contains three important catalytic residues (Asp363, Glu399, and Asp471) situated at the bottom of the active site cleft. The dimer active site exhibits a buried interface of 1523 Å, and a total of 6 subsites, where subsite 1 contains the NYWDYDP motif important which facilitates substrate interactions of the glucose rings. One of the interesting features which separate TreX from similar isoamylases and pullulanases, it the presence of a helix α4 loop (aa 228-238) situated at the bottom of the substrate binding groove. It is suggested that this helix α4 loop may provide a stable binding region for branched substrates with long chains, and therefore increase the activity of this GDE [1].
Upon tetramerization, TreX’s active site undergoes a substantial change in conformation, and thus displays both structural and functional differences when compared to the TreX dimer and even to other GDEs. Studies have revealed that the TreX tetramer displays a 4-fold increase in catalytic activity, when compared to the dimer. It is postulated that the conformational change in the active site only occurs in the presence of branched substrates with long chains, such as glycogen, because it establishes an ideal binding site for these types of substrates. To illustrate, the TreX tetramer is composed of two dimers, whose active sites face each other with a slight offset. As a consequence, particular regions of one dimer, such as lid 1(aa 92-97) and lid 2 (aa 315-322), are situated in the active site of the other dimer. This arrangement of the dimers’ structural lids results in the formation of a channel like cavity and a conformational change in a loop (aa 399 – 416) situated within the active site. The two lid structures have been greatly associated with increased α-1,4-transferase activity in TreX, and interestingly, their conformation exhibits structural similarity to that of other glucosyltransferase enzymes, which also encompass structural lids close to their active sites. Thus, the implicated function of the lid structures during catalysis is to interact with acceptor molecules, such as glycogen, and provide stability to the complex, so that glycogen may be broken down into long maltooligosacchardies [1].
Mechanism
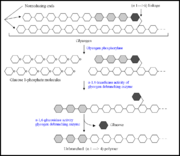
Figure 1. Diagram illustrating the breakdown of glycogen near an (α1→6) branch point, and the steps where the α-1,6-glycosidase and the α-1,4-transferase activity of the glycogen debranching enzyme takes place. Diagram adapted from
[6].
In human metabolism, glycogen breakdown involves several enzymes, two of which are glycogen phosphorylase and glycogen-debranching enzyme [6]. Glycogen phosphorylase is responsible for the successive removal of glucose 1-phosphate molecules at the non reducing ends of glycogen branches [6]. However, this enzyme’s activity ceases when it has reached a point four glucose residues away from an (α1→6) branch point. Upon this, the GDE takes over and catalyzes the transfer of three branched glucose units to the nonreducing end of another branch to yield an (α1→4) linkage. The α-1,6-glycosidase activity of the GDE liberates the non-transferred glucose unit involved in a (α1→6) bond [6].
Although TreX is structurally different from the yeast and human GDE, it does however share functional similarities, and therefore a general human GDE mechanism is shown. In both species the GDE catalyzes an intermolecular transfer of glucose polymers from one 1,4-α-D-glucan branch to another 1,4-α-D-glucan branch nearby, In addition, both enzymes carry out the hydrolysis of an α-1,6-glycosidic linkage [7]. TreX does however show high specificity for side chains which are composed of 6 or more glucose residues [5].
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 Woo EJ, Lee S, Cha H, Park JT, Yoon SM, Song HN, Park KH. Structural insight into the bifunctional mechanism of the glycogen-debranching enzyme TreX from the archaeon Sulfolobus solfataricus. J Biol Chem. 2008 Oct 17;283(42):28641-8. Epub 2008 Aug 14. PMID:18703518 doi:10.1074/jbc.M802560200
- ↑ 2.0 2.1 Stam MR, Danchin EG, Rancurel C, Coutinho PM, Henrissat B. Dividing the large glycoside hydrolase family 13 into subfamilies: towards improved functional annotations of alpha-amylase-related proteins. Protein Eng Des Sel. 2006 Dec;19(12):555-62. Epub 2006 Nov 2. PMID:17085431 doi:10.1093/protein/gzl044
- ↑ Shim JH, Park JT, Hong JS, Kim KW, Kim MJ, Auh JH, Kim YW, Park CS, Boos W, Kim JW, Park KH. Role of maltogenic amylase and pullulanase in maltodextrin and glycogen metabolism of Bacillus subtilis 168. J Bacteriol. 2009 Aug;191(15):4835-44. Epub 2009 May 22. PMID:19465663 doi:10.1128/JB.00176-09
- ↑ Kubo A, Fujita N, Harada K, Matsuda T, Satoh H, Nakamura Y. The starch-debranching enzymes isoamylase and pullulanase are both involved in amylopectin biosynthesis in rice endosperm Plant Physiol. 1999 Oct;121(2):399-410. PMID:10517831
- ↑ 5.0 5.1 Park JT, Park HS, Kang HK, Hong JS, Cha H, Woo EJ, Kim JW, Kim MJ, Boos W, Lee S, Park KH (2008). "Oligomeric and functional properties of a debranching enzyme (TreX) from the archaeon Sulfobus solfataricus P2.". Biocatalysis and Biotransformation 26: 76–85.
- ↑ 6.0 6.1 6.2 6.3 Nelson, D. and Cox, M. Lehninger Principles of Biochemistry (5th Ed.), W.H. Freeman and Company, New York (2008).
- ↑ Park HS, Park JT, Kang HK, Cha H, Kim DS, Kim JW, Park KH. TreX from Sulfolobus solfataricus ATCC 35092 displays isoamylase and 4-alpha-glucanotransferase activities. Biosci Biotechnol Biochem. 2007 May;71(5):1348-52. Epub 2007 May 7. PMID:17485831


