From Proteopedia
proteopedia linkproteopedia link Introduction
|
|
| 1js3, resolution 2.25Å ()
|
| Ligands:
| , ,
|
| Activity:
| Aromatic-L-amino-acid decarboxylase, with EC number 4.1.1.28
|
| Related:
| 1js6
|
| Structural annotation:
|
| Resources:
| CATH : 1Js3A01, 1Js3A03, 1Js3A02, 1Js3B03, 1Js3B02, 1Js3B01
InterPro : Ipr010977, Ipr002129, Ipr015421, Ipr015424, Ipr015422
Pfam : PF00282
SCOP : d1js3a_, d1js3b_
UniProt : P80041
|
|
|
|
|
|
| Resources:
| FirstGlance, OCA, RCSB, PDBsum
|
| Coordinates:
| save as pdb, mmCIF, xml
|
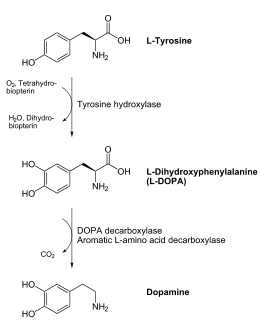
DOPA decarboxylase (DDC, aromatic L-amino acid decarboxylase, tryptophan decarboxylase, 5-hydroxytryptophan decarboxylase, AAAD) is an approximately 52 kDa protein that belongs to the aspartate aminotransferase family (fold type 1) of PLP-dependent enzymes. The catalytically active form of the enzyme exists as a homodimer, typical of this class of enzymes.[1] The homodimeric form of the enzyme purified from sus scrofa is shown in complex with the inhibitor carbidopa to the right. DOPA decarboxylase is responsible for the synthesis of dopamine and serotonin from L-DOPA and L-5;hydroxytryptophan, respectively. Due to its role in neurotransmitter synthesis, DOPA decarboxylase has been implicated in Parkinson's disease, a disease thought to be the result of the degeneration of dopamine-producing cells in the brain. Currently, treatment for the disease is aimed at DOPA decarboxylase inhibition, which would allow greater amounts of exogenously administered L-DOPA to reach the brain. [2]
Structure
DOPA decarboxylase is a homodimeric enzyme, with each monomer composed of three distinct domains. The large domain contains the PLP binding site, and consists of a seven stranded mixed beta-sheet that is surrounded by eight alpha-helices, resulting in a typical alpha/beta fold. The C-terminal domain is comprised of a four-stranded anti-parallel beta-sheet that has three alpha-helices packed against the face opposite to the large domain. Although the aforementioned domains exist in all members of this family of PLP-dependent enzymes, including bacterial ornithine decarboxylase (OrnDC) and dialkylglycine decarboxylase (DGD), the is unique to DOPA decarboxylase, and is a representative case of domain swapping. This domain is composed of two parallel helices linked by an extended strand, which essentially lies like a flap over the second subunit. The N-terminal domain of one monomer packs on top of the other monomer, resulting in an extended dimer interface, and thus it is most likely stable only in the dimeric form of the enzyme.
References
- ↑ Schneider G, Kack H, Lindqvist Y. The manifold of vitamin B6 dependent enzymes. Structure. 2000 Jan 15;8(1):R1-6. PMID:10673430
- ↑ Burkhard P, Dominici P, Borri-Voltattorni C, Jansonius JN, Malashkevich VN. Structural insight into Parkinson's disease treatment from drug-inhibited DOPA decarboxylase. Nat Struct Biol. 2001 Nov;8(11):963-7. PMID:11685243 doi:http://dx.doi.org/10.1038/nsb1101-963