This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox SouthUniversity1
From Proteopedia
| |||||||||||
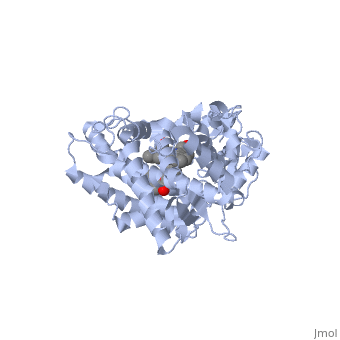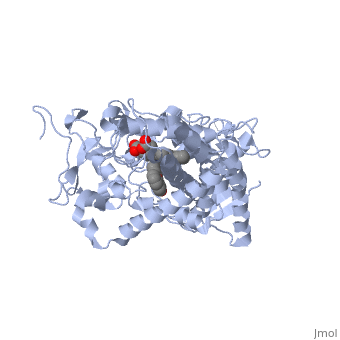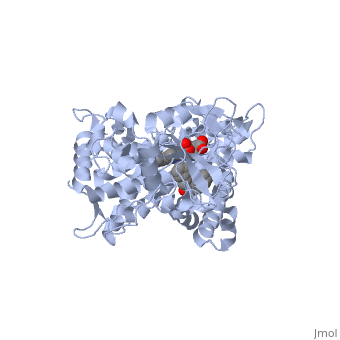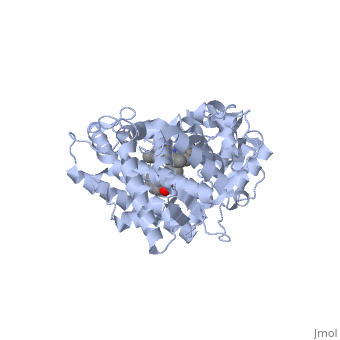
Different members of the CYP450 enzymes preferentially metabolize different xenobiotics. What features of a drug do you think might cause it to be metabolized by one CYP versus another? Draft: <illustration of residues in active site> Draft animation Draft animation
draft: <insert quiz here on electrostatic and steric factors>
draft: <insert animations here contrasting the shape of the active site of CYP3A4 and e.g. CYP2E1>
draft: <insert examples of substrates for 2 different CYPs, overlay them on top of each other illustrate the molecular commonalities>
draft <illustrate the size of the binding pockets and their electronic character>
draft test of scene names:
<Draft Quiz>
References The structure of CYP1A2 shown above is found in the Research Collaboration of Structural Bioinformatics Protein Database as entry 2HI4.