1gpa
From Proteopedia
| |||||||||
| 1gpa, resolution 2.90Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||||
| Non-Standard Residues: | |||||||||
| Activity: | Phosphorylase, with EC number 2.4.1.1 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
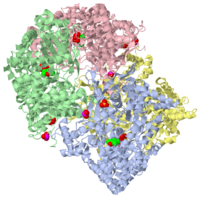
STRUCTURAL MECHANISM FOR GLYCOGEN PHOSPHORYLASE CONTROL BY PHOSPHORYLATION AND AMP
Template:ABSTRACT PUBMED 1900534
About this Structure
1gpa is a 4 chain structure with sequence from Oryctolagus cuniculus. The December 2001 RCSB PDB Molecule of the Month feature on Glycogen Phosphorylase by David S. Goodsell is 10.2210/rcsb_pdb/mom_2001_12. Full crystallographic information is available from OCA.
See Also
Reference
- Barford D, Hu SH, Johnson LN. Structural mechanism for glycogen phosphorylase control by phosphorylation and AMP. J Mol Biol. 1991 Mar 5;218(1):233-60. PMID:1900534