This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
7rsa
From Proteopedia
| |||||||||
| 7rsa, resolution 1.26Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||||
| Activity: | Pancreatic ribonuclease, with EC number 3.1.27.5 | ||||||||
| |||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
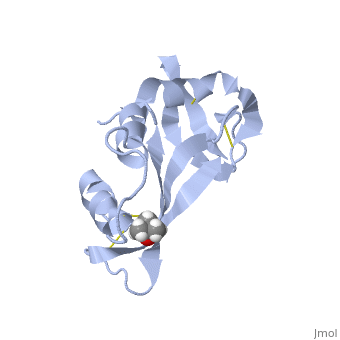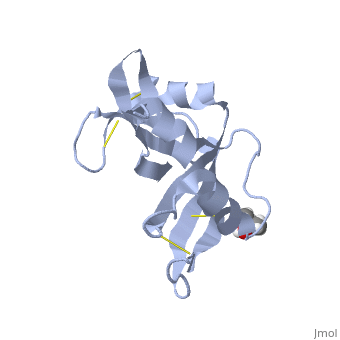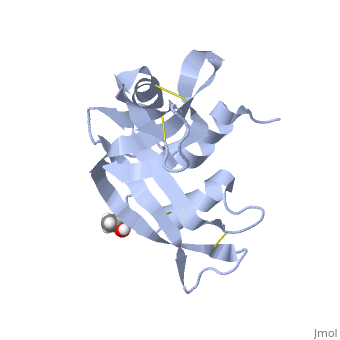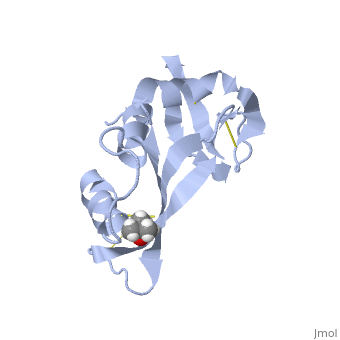
STRUCTURE OF PHOSPHATE-FREE RIBONUCLEASE A REFINED AT 1.26 ANGSTROMS
Template:ABSTRACT PUBMED 3401445
About this Structure
7rsa is a 1 chain structure with sequence from Bos taurus. Full crystallographic information is available from OCA.
See Also
Reference
- Wlodawer A, Svensson LA, Sjolin L, Gilliland GL. Structure of phosphate-free ribonuclease A refined at 1.26 A. Biochemistry. 1988 Apr 19;27(8):2705-17. PMID:3401445
- Kobe B, Deisenhofer J. Mechanism of ribonuclease inhibition by ribonuclease inhibitor protein based on the crystal structure of its complex with ribonuclease A. J Mol Biol. 1996 Dec 20;264(5):1028-43. PMID:9000628 doi:http://dx.doi.org/10.1006/jmbi.1996.0694
- Richardson JS, Richardson DC. Natural beta-sheet proteins use negative design to avoid edge-to-edge aggregation. Proc Natl Acad Sci U S A. 2002 Mar 5;99(5):2754-9. PMID:11880627 doi:10.1073/pnas.052706099