General Function
1diz Escherichia coli 3 methyladenine DNA glycosylase II (AlkA) is a DNA repair enzyme that initiates base excision repair for the removal of alkylated bases. There are many environmental toxins and cellular agents that may alkylate DNA bases. One example is Aflatoxin, a compound that is converted into a reactive epoxide by cytochrome P450 and attacks guanosine at its N-7 atom to form an alkylated base.[1] The alkylated DNA bases prevent regulatory proteins from binding to DNA and blocks replicative polymerases to stop DNA synthesis.[2] AlkA initiates base excision repair by first locating and binding to the alkylated DNA. It has broad substrate specificity and is able to remove a variety of different alkylated bases. AlkA then flips the affected base out of the DNA double helix and into the active site of the enzyme. Once in the active site, AlkA hydrolyzes the glycosidic bond to release the damaged base and leave the sugar phosphate backbone intact. This creates the AP site that is either devoid of a purine or pyridine. The AP site signals to other base excision repair enzymes to insert an undamaged nucleotide based on the undamaged complementary strand and seal the DNA.[1]
Key Structures
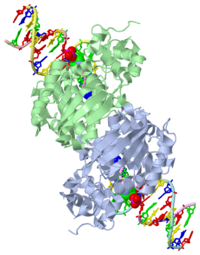
AlkA is composed of three main domains with dimensions of approximately 50 Angstroms, 45 Angstroms, and 25 Angstroms.[3] The first domain (residues 1-112) is the that is composed of a five stranded antiparallel beta sheet and two alpha helices.[3] The (residues 113-230) contains seven alpha helices that create a hydrophobic core.[3] The third domain (resudues 231-282) is the that contains a bundle of three alpha helices. [3] The three domains of AlkA create a distinct binding structure that is lined with electron rich aromatic residues to form a hydrophobic cleft. The cleft is found between domains 2 and 3, and forms the binding pocket for the alkylated base. The hydrophobic cleft has the ability to widen between domains 2 and 3 for accepting a variety of alkylated bases.[3]
AlkA is a member of the helix-hairpin-helix (HhH) family of DNA glycosylases, where two compact alpha helical structures are connected by a hairpin loop.[4] In AlkA, the is located on the rim of the active site and is composed of residues 202-227.[2] The HhH sequence is responsible for binding the damaged DNA by van der Waals interactions, a few hydrogen bonds, and metal ion interactions.
Active Sites and Mechanism
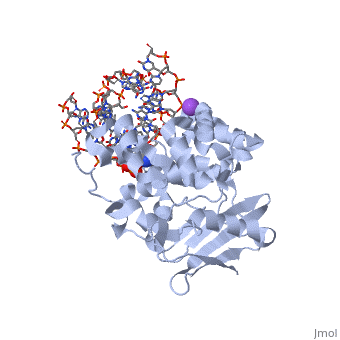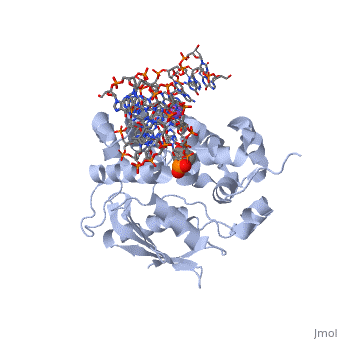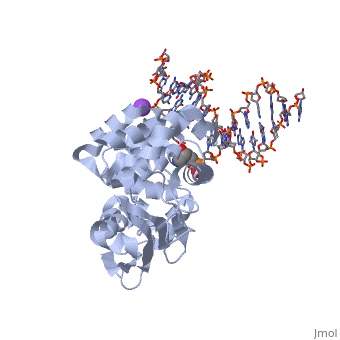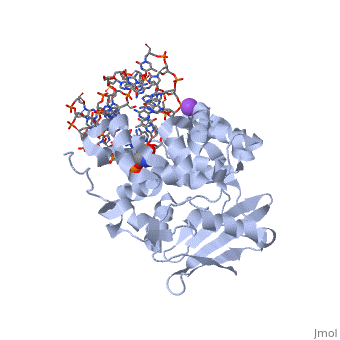
The HhH segment of AlkA connects to the phosphodiester backbone of DNA by hydrogen bonding and . The binding of the HhH segment to DNA stabilizes the damaged base and creates a 66 degree bend away from the protein that widens the minor groove of DNA.[4] fits into the minor groove between base pairs and allows the alkylated base to be flipped into the active site.[2]
The AlkA active site is located where the second and third domains are separated by a deep nonpolar cleft that is lined with the aromatic side chains Phe18, Tyr273, Trp272, Tyr222, and Trp218.[3] These side chains create a for the alkylated base.[3] Once the damaged base is in the active site, stabilizes the flipped out alkylated base in the binding pocket by aromatic ring-stacking interactions.[4] is essential for allowing the reaction to proceed, and points into the nonpolar pocket in order to allow stabilization of a carbocation intermediate. This stabilization is what allows the cleavage of the glycosidic bond on the damaged base.[4]
DNA Interaction
The AlkA protein stabilizes DNA by polar and nonpolar interactions with the DNA backbone. There are few positively charged residues on the AlkA’s binding surface that could interact with the DNA backbone, which is why the majority of polar interactions involve the DNA strand containing the alkylated base.
DNA is held in place by interactions with the HhH segment, which acts as a platform to create the 66 degree bend in DNA mentioned earlier. The bending of the DNA relieves tensional strain created by the widening of the minor groove for base flipping.[2] Amide nitrogens on residues Gly214, Gly216, and Thr219 with DNA’s phosphodiester backbone, while residues Gln210, Phe212, and Ile215 the sodium metal ion that interacts with DNA further stabilize it in the molecule.[2]
In order to keep the DNA strand bound to the protein, AlkA depends on van der Waals interactions on the minor groove of DNA. The van der Waals interactions play a major role in AlkA’s preference for double stranded DNA.[2] An example of this may be seen with the Pro175 wedged into the minor groove of DNA and anchoring it by van der Waals interctions.[2]
The essential Asp238 residue creates hydrogen bonds with the alkylated DNA base that has been flipped into the active site. Its high affinity binding for the modified base holds it in the active site for cleavage of the glycosidic bond. An inhibitor or mutation to the Asp238 residue would ultimately prevent the cell's ability to modify DNA by base excision repair.