This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 933
From Proteopedia
| This Sandbox is Reserved from 01/04/2014, through 30/06/2014 for use in the course "510042. Protein structure, function and folding" taught by Prof Adrian Goldman, Tommi Kajander, Taru Meri, Konstantin Kogan and Juho Kellosalo at the University of Helsinki. This reservation includes Sandbox Reserved 923 through Sandbox Reserved 947. |
To get started:
More help: Help:Editing |
Contents |
DNA Binding Domain of LEAFY: A Master Regulator in Flower Development
Introduction
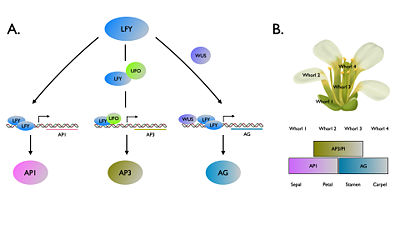
FLORICAULA/LEAFY (FLO/LFY) genes encode a plant specific transcription factor family that controlling floral fate of reproductive phase. (ref 1) In the plant model system Arabidopsis thaliana , ‘’LFY’’ also acts upstream of floral homeotic genes to modulate organ identity. (ref 2) LFY activates the organ identity genes by binding to promoter regions of floral organ identity genes. LFY can directly bind to the promoter to APELATA1 (AP1), while co-regulators UNUSUAL FLORAL ORGANS (UFO) (ref 3) and WUSCHEL (WUS) (ref 4) are required for increment of binding affinity to promoter regions of APELATA3 (AP3) and AGAMOUS (AG), respectively. The exact mechanism how LFY binds to these promoters has yet to be well elucidated until the first structure report about LFY-pAP1 and LFY-pAG (ref 5). Among land plants, FLO/LFY homologs share a highly conserved DNA binding region that a hypothesis claimed substitution in this domain might result in the functional divergence (ref 6). Recently, a new study (ref 7) provided new insights of structural basis of evolution by changing DNA binding.
Structural Basis of LEAFY binding
|
General information about the structure
DNA recognition is conducted by a HTH-like motif
DNA binding required cooperative dimerization
LEAFY Evolution
Reference
1 Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM (1992). "LEAFY controls floral meristem identity in Arabidopsis". Cell 69 (5): 843–859. doi:10.1016/0092-8674(92)90295-N. PMID 1350515. 2