This is a default text for your page Sydney Pate/Sandbox 1. Click above on edit this page to modify. Be careful with the < and > signs.
You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
Structure
[3]
[4]
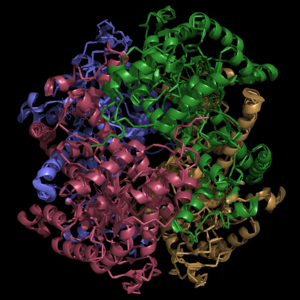
The active site of isocitrate lyase consists of two catalytic residues: Cys191 and His193. Additionally, there are several other amino acid side chains present that form hydrogen bonding opportunities with isocitrate to catalyze the breakdown reaction to glyoxylate and succinate. Ser91, Gly92, Trp93, and Arg228.
Function
Disease
Relevance
Structural highlights
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.