This is a default text for your page '. Click above on edit this page' to modify. Be careful with the < and > signs.
You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
Biological Function
Structural Overview
CzrA functions as a dimer. The dimerize at the czr operon, repressing gene transcription. Each monomeric unit contains seen in purple and displayed in yellow. While the function of the beta sheets are not yet known, key helices regulate the binding of DNA and Zn +2 . The is the location of DNA binding and the contains the Zn +2 binding site. As Zn +2 binds, the alpha 4 helices are , repressing their DNA binding ability.
Binding of DNA
The have been found to occur at the Ser 54 and 57 along with His 58. These residues are likely to interact with the 5'-TGAA sequence found in the half-site of the DNA. These residues are found in the N terminal of the R helix. The residues involved in the are Val 42 and Gln 53. This was experimentally determined by replacing the Gln and Val with Ala residues and measuring the binding capacity; In a previously published article
[3], the DNA bound state of CzrA was tested by using the known critical residues for DNA interactions. Critical residues, Gln53, Val42, Ser54, Ser57, and His58, were replaced with Ala and then compared to the kinetics of the wild type protein. Replacing only the Q53 and V42 residues resulted in an 11-fold and 160-fold decrease in K
a, respectively. Other residues such as S54, S57, and H58 were also replaced with Ala residues, and it was found that these mutations caused binding similar to the fully inhibited Zn
2+ bound state. Table 1 in this same article shows the different K
observed, and the measured decrease in K
observed for each mutation. The bind between the DNA and the protein can be attributed to losing certain intermolecular forces such as possible hydrogen bonding when changing from Gln and Ala, and a loss of London Dispersion forces in the Val to Ala change.
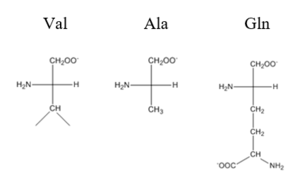
Comparison of Val, Ala, and Gln residues
The differences in binding favorability can also be seen when comparing the ΔG for the Apo-state vs. the DNA bound state and the Zinc vs. the Zinc and DNA bound state. These ΔGs were found to be -15.2kcal/mol and -9kcal/mol respectively[4]. This agrees with previously published data showing the Zinc binding inhibits the affinity the protein has to DNA.
Zinc Binding
Zinc is an allosteric inhibitor to CzrA. Two Zn +2 ions may bind to the dimer, at the location of the helix from each monomer. As zinc binds, the alpha 5 helices to inhibit the DNA binding residues. Furthermore, CzrA must be in its dimer form for zinc to bind. The is formed by two residues from each monomer, so Zn+2 cannot bind to the monomer. The is formed by Asp84 and His86 from one monomer, and His97 and His100 from the other monomer.
The zinc+2 ion forms a tetrahedral complex with the four residues (Figure 1). This allows other metal ions to act as allosteric inhibitors to CzrA. Any metal that may form a tetrahedral complex will have some affinity for CzrA, assuming it is not too large to fit into the pocket. However, the metal binding pocket of CzrA has been optimized to bind Zn+2 with the highest affinity. As CzrA is a transcriptional repressor, binding of Zn+2 to the dimer will activate the czr operon. Zn+2 is preferred as CzrB opens a Zn+2 channel, allowing the excess zinc ions to export the cell.
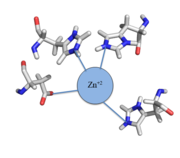
Figure 1:Zn
+2 tetrahedral binding complex
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ doi: https://dx.doi.org/10.1073pnas.0905558106
- ↑ Chakravorty DK, Wang B, Lee CW, Giedroc DP, Merz KM Jr. Simulations of allosteric motions in the zinc sensor CzrA. J Am Chem Soc. 2012 Feb 22;134(7):3367-76. doi: 10.1021/ja208047b. Epub 2011 Nov , 14. PMID:22007899 doi:http://dx.doi.org/10.1021/ja208047b


