This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Alexis Neyman/Sandbox 1
From Proteopedia
All About SRp20
SRp20 Structure
Anything in this section will appear adjacent to the 3D structure and will be scrollable.
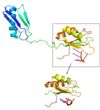
BackgroundProtein of interest: SRp20 Significance: RNA binding protein Organism: Homo Sapiens, Streptococcus sp. Group G Characteristics: Ser/Arg-rich domain Basic function: Splicing Factor 164 aa: half of it belongs to the RS (Arg/Ser) domain MW: 19 kDA Determined via HSQC NMR (heteronuclear single quantum coherence) FunctionGeneral Funtions (GO INTO DETAIL ABOUT EACH)RNA processing Termination of transcription Protein translation Cellular proliferation/maturation Involved in Insulin signaling (loss in liver results in wrong splicing of glucose metabolism genes) Embryogenesis Splicing FactorThe spliceosome is a complex responsible for splicing pre-mRNA to mature mRNA, composed of snRNA and associated proteins Introns are spliced out, while exons are ligated together Splicing is necessary for mRNA maturation, enabling it to exit the nucleus Alternative SplicingAlternative splicing allows many different proteins to be produced from one mRNA by splicing different combinations of introns and exons Changes or defects in this process linked to human diseases RNA processing factors may be targets for future therapies SR ProteinsCharacteristics: One or two N’ term RRMs (RNA recognition motifs) Arg/ser-rich domain downstream Most serine residues phosphorylated The N’ term RRM provides substrate specificity The C’ term RS domain aids protein-protein interactions (simplifying spliceosome recruitment) RMMN-terminal RNA recognition motif standard for an RRM A four stranded β-sheet and two α-helices Most commonly, three aromatic side-chains (in β-3 and β-1 strands) accommodate two nucleotides Recognition enables binding of SRp20 to RNA mRNA MaturationRelationship to other proteinsRelationship to 9G8SRp20 and 9G8 are both sequence specific RNA binding proteins. They are the smallest members of the Serine-and-Arginine Rich (SR) protein family. Both RNA Recognition Motifs (RRMs) have a similar βαββαβ topology. SRp20 and 9G8 are 80% identical. The sequence alignment shows the alignment of the RRMs of SRp20 and 9G8 (Hargous et al., 2006). SRp20 binds pyrimidine rich areas while 9G8 binds purine rich areas.This difference in binding comes from the fact that 9G8 has a zinc knuckle that recognizes GAC triplets (Cavaloc et al., 1999). 9G8s RRM is followed by a zinc knuckle and then the SR domain whereas SRp20s RRM is followed directly by the SR domain. When 9G8 lacks a zinc knuckle, it binds pyrimidine-rich sequences like SRp20 (Hargous et al., 2006). The zinc knuckle of 9G8 contains glycine residues at positions 5 and 8 and charged residues at positions 6 and 13 that are highly conserved (Cavaloc et al., 1999). Due to the poor solubility problem, a structure for the zinc knuckle of 9G8 is not available to show in an image. A study by Huang and Steitz showed that 9G8 and SRp20 promote the export of mRNA from the nucleus. There is a 101-nt sequence in the coding region of mouse histone H2a mRNA that promotes the the export of intronless human β-globin cDNA transcripts (Huang and Carmichael, 1997). Of this 101-nt sequence, there is specifically a 22-nt sequence that is necessary for export activity. When this 101-nt sequence was not present, mRNA was not properly processed, and RNA accumulated in the nucleus; however, when the 101-nt sequence was present mRNA export from the nucleus was improved almost 4-fold. This 101-nt sequence improves export and polyadenylation, but removing the 22-nt sequence stopped both of these activities suggesting that the 22-nt sequence is required for successful nuclear export of β-globin cDNA. UV cross-linking and immunoprecipitation experiments determined the proteins that specifically associate with the 22-nt sequence. It was determined that SR proteins were associating with the 22-nt sequence by adding antibodies specific for SRp20 then 9G8. SRp20 antibodies inhibited mRNA export 3-fold while 9G8 antibodies inhibited mRNA export at least 10-fold showing that SRp20 and 9G8 are active factors that promote mRNA export. It was shown that SRp20 and 9G8 are cross-linked to polyadenylated RNA in the nucleus and cytoplasm showing that both proteins play a direct role in mRNA export from the nucleus (Huang and Steitz, 2001). DiseaseCancerOncogenic signaling pathways such as the WnT pathway cause-effect relationship with increase expression of SRp20 Downregulation of SRp20 promotes cellular senescence through AS of TP53 and generating p53 (tumor suppressor gene) Silencing SRp20 in cancer cells: slower cell proliferation Activator of the AS of gene CD44 Pyruvate Kinase M (PK-M gene) Regulate Rac1b protein expression in colorectal tumor cells Alzheimer'sTRKB: AS results in TrkB-Shc transcript Human Papillomavirus: HPVIncreases SRp20 which in turn regulates the gene expression of HPV via interaction with A/C-rich RNA nucleobases Overexpression of it involved in HPV-induced cancers such as cervical cancer Structural highlightsPoor Solubility ProblemThe SRp20 protein has poor solubility in its free state. This made it impossible to determine the structure of SRp20 using HSQC Spectroscopy without a modification to the free state protein. This problem was resolved by studying the proteins after fusing the RRM (RNA-recognition motif) with the immunoglobulin G-binding domain 1 of Streptococcal Protein G GB1 solubility tag.
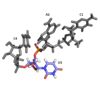
RNA Interactions1H-15N HSQC results showed a large hydrophobic β-sheet on the RRM binds to the RNA with all four bases contacting one of the four aromatic residues” (hydrophobic interactions) Other structural studies show that amino acids of the β-hairpin are directly hydrogen bonded to bases of nucleic acid targets Using a smaller peptide chain reduced the NMR broadening seen with longer peptides (allowing for structure determination), though the binding affinity was also reduced C1 and A2 stack on Y13 in β1 and F50 in β3 (aromatic side chains), respectively.
The residue F48 inserts between the sugar rings of C1 and A2 Looking at the ligandThe conformation of U3 and C4 is unusual because U3 bulges out while C4 stacks over A2, partially.
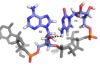
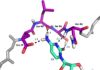
also adopts an unusual syn conformation C4 is maintained in its position by a RRM Domain InteractionsU3 interacts with and with the β2-3 loop of the RRM
C1 amino protons hydrogen bond with Leu 80 carbonyl oxygen and Glu 79 side-chain carbonyl oxygen. C1 N3 hydrogen bonds with Asn 82 amide. C1 O2 hydrogen bonds with Ser 81 hydroxyl group. Ser-Arg Rich DomainSpecificity4 nt can be accommodated by RRM β-sheet, but recognition is only partially sequence specific. CAUC C1 more specific, A2 and U3 less specific It is uncertain whether C4 is specifically recognized by the RRM A is prefered over G at the 2 position, but no indication of preference over U or C U3 is even less specific, could be C, G or A The recognition of C1 is functionally necessary because a C to G mutation within the histone mRNA can impair RNA export Advantages of low specificityLess evolutionary pressure on bound RNA (ideal for exonic sequences) More RNA sequences can be targeted SRp20 can associate/disassociate with RNA more easily Important for highly dynamic RNA metabolism processes RNA binding affinity can be modulated by protein-protein interactions (which are dependant on the level of phosphorylation) This can be used to tune post-transcriptional gene expression Relevance and ConclusionsUnderstanding and recognizing the mechanisms that SRp20 is involved in can help find treatment and management of cancer patients The use of SR proteins (such as SRp20) may in the future be used for targeted therapy No structure for the C-term domain Future DirectionsRecent structural studies emphasize that not only the β-sheet surface but also the loops connecting β-strands and a-helices can be crucial for nucleic acid recognition → future research looking more closely at this possibility Farther research could be done investigating the process of RS domain phosphorylation and how it controls splicing Look farther into the specificity or lack of specificity relating to SR proteins Mutate specific residues in order to map out SRp20 mechanistic function
ReferencesCle’ry A, Blatter M, Allain FHT. 2008. RNA recognition motifs: boring? Not quite. Current Opinion in Structural Biology 18: 290–298. Doi: 10.1016/j.sbi.2008.04.002 Corbo C, Orrù S, Salvatore F. 2013. SRp20: An overview of its role in human diseases. Biochemical and Biophysical Research Communications 436: 1–5. Doi: 10.1016/j.bbrc.2013.05.027 Hargous Y, Hautbergue GM, Tintaru AM, Skrisovska L, Golovanov AP, et. al. 2006. The EMBO Journal ) 25: 5126–5137. Doi: :10.1038/ sj.emboj.7601385
| ||||||||||||