Nuclear polyadenylated RNA-binding protein
From Proteopedia
Contents |
Introduction
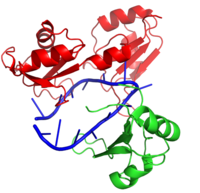
Hrp1 is a polyadenylation factor found in Saccharomyces cervisiae (yeast) [1]. This protein recognizes and binds to an RNA sequence in the 3'UTR of the messenger RNA (mRNA) upstream from the cleavage site called the polyadenylation enhancement element (PEE) [1]. Upon binding to the RNA, Hrp1 helps recruit additional proteins necessary for the cleavage and polyadenylation of the RNA molecule [1]. The unique structural features of the Hrp1-PEE complex reveals the mechanism by which Hrp1 is able to recognize and bind to its specific RNA sequence at the atomic level [1].
Structure
| |||||||||||
Interaction with RNA15
RNA15 is another RNA-binding protein with a single N-terminal RNA recognition motif (RRM) [2]. RNA15 recognizes an A-rich positioning element (PE) downstream from the PEE but upstream from the 3' cleavage site [2]. The recognition of the PE by RNA15 is crucial for precise cleavage of the RNA molecule. Hrp1 and RNA15 are held together by a separate protein, RNA14 [2]. These proteins act together to anchor the polyadenylation and cleavage protein machinery relative to the cleavage site for precise 3'-end processing [2].
Relationship to other proteins
The RNP-type RBD is found in many proteins involved in post-transcriptional pre-mRNA processing (5'-end capping, splicing, 3'-end cleavage and polyadenylation, and transport from the nucleus)[3]. The unique RBD of Hrp1 enables the protein to bind an RNA sequence that differs in both length and content from the RNA sequences of other RNA-binding and mRNA processing proteins such as sex lethal, Poly (A)-binding protein (PABP), and HuD [1]. Like Hrp1, each of these proteins belong to the class of single strand proteins composed of two canonical RBDs; however, these proteins are differentiated by their target RNA sequence, their interactions with RNA at the atomic level, and their interdomain contacts [1]. One way in which Hrp1 differentiates itself from these other proteins is by the fact that Hud, sex lethal, and PABP all contain at least one intra-RNA base-base stacking interaction, a feature that is not found in the Hrp1-PEE complex [1]. It is possible that the intra-RNA interactions found in these other proteins is replaced by the crucial Trp168-Ade4 stacking interaction found in the Hrp1 complex [1]. The fact that the intra-RNA base-base stacking interactions are replaced by the Trp168-Ade4 in the Hrp1-PEE complex might also explain why the Hrp1-RNA interface involves only 6 nucleotides whereas PABP, sex lethal, and HuD require a longer 8-10 nucleotide sequence in the RNA binding pocket [1].
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 Perez-Canadillas JM. Grabbing the message: structural basis of mRNA 3'UTR recognition by Hrp1. EMBO J. 2006 Jul 12;25(13):3167-78. Epub 2006 Jun 22. PMID:16794580
- ↑ 2.0 2.1 2.2 2.3 Leeper TC, Qu X, Lu C, Moore C, Varani G. Novel protein-protein contacts facilitate mRNA 3'-processing signal recognition by Rna15 and Hrp1. J Mol Biol. 2010 Aug 20;401(3):334-49. Epub 2010 Jun 19. PMID:20600122 doi:10.1016/j.jmb.2010.06.032
- ↑ Clery A, Blatter M, Allain FH. RNA recognition motifs: boring? Not quite. Curr Opin Struct Biol. 2008 Jun;18(3):290-8. doi: 10.1016/j.sbi.2008.04.002. PMID:18515081 doi:http://dx.doi.org/10.1016/j.sbi.2008.04.002
Proteopedia Page Contributors and Editors (what is this?)
Cory A. Wuerch, Matthew Douglas Moore, Savannah Davis, Michal Harel, Jaime Prilusky