User:Courtney Brown/Sandbox 1
From Proteopedia
The Human Histone H3/K9 Deacetylase, HDAC8
This is a default text for your page Courtney Brown/Sandbox 1. Click above on edit this page to modify. Be careful with the < and > signs. You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
IntroductionHistonesHistones are a family of basic, positively charged proteins that associate with DNA inside the nucleus to help condense the DNA into chromatin [3]. The nuclear DNA is wrapped around the histone in order to fit in the nucleus. Nucleosomes are chromatin beads made up of DNA wrapped around eight histone proteins, or a histone octamer [3]. The modification of histones are a type of epigenetics, where changes are made in gene expression without altering the DNA sequence. Four different examples of modifying histones including histone acetylation, histone deacetylation, histone methylation and histone demethylation [3]. Histone Deacetylases (HDACs)ε-Amino-lysine acetylation is a type of histone modification that controls the stability of proteins and biological function in eukaryotic cells [4]. Histone Deacetylation is the reversal process for this acetylation modification. There are different classes of HDACs based on phylogenetic analysis: •Class I - HDACs 1-3 and 8, which are homologous to yeast Rpd3 •Class II - HDACs 4-7, 9 and 10, which are homologous to yeast Hda1 •Class III - Sirtuin deacetylases •Class IV - HDAC 11 [4]. HDACs 1-11 are metalloenzymes and require a zinc ion for deacetylation [4]. HDAC8Histone Deacetylase 8 is an enzyme found in Homo sapiens. HDAC8 is 388 residues long and consists of eight-stranded parallel β-sheets surrounded by 11 α-helices [4]. HDAC8 is the only functional HDAC that is found to be a single polypeptide instead of being high-molecular-weight multi-protein complexes [4]. The substrate bound to the HDAC8 includes an acetyl group, one arginine, one histidine, two lysines and MCM, a Coumarin fluorescence tag. StructureGeneral Structure Information
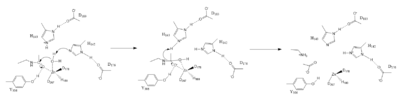
InhibitorPotassium Binding SiteDeacetylationActive SiteSubstrates are deacetylated via a classic 2+ metal ion mechanism. The Zinc ion by withdrawing electron density from the water due to the positive charge, making the water more acidic, therefore a better nucleophile. The Zinc ion also coordinates to the carbonyl oxygen of the acetyl group on the lysine, polarizing the carbonyl carbon, making it more electrophilic. deprotinates the water, the first step of the deacetylation. His 142 is closer to the water molecule than His 143, which was also thought to deprotinate the water. However, a mutation done to His 143 only reduced activity, not abolished it, showing it is important but not crucial. His 143 is thought to orient the substrate by protinating the amine group on the leaving group of the reaction [4].
DiseaseReferences
Student Contributors
| ||||||||||||