This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Epoxidase
From Proteopedia
| |||||||||||
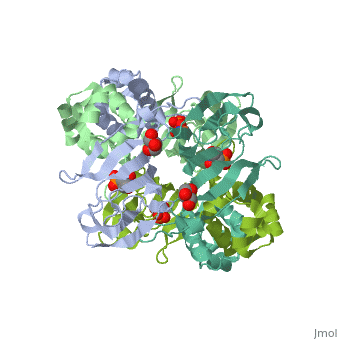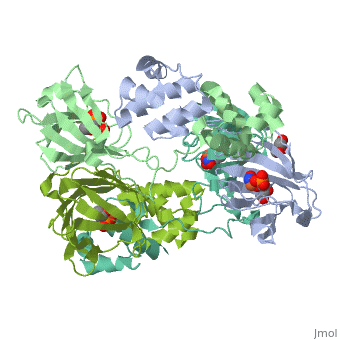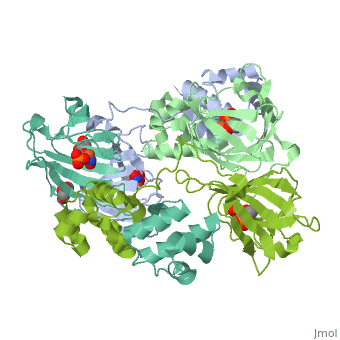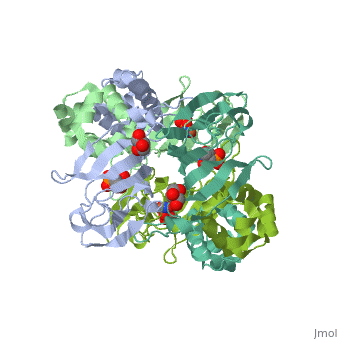
3D structures of epoxidase
Updated on 23-June-2019
References
- ↑ McLuskey K, Cameron S, Hammerschmidt F, Hunter WN. Structure and reactivity of hydroxypropylphosphonic acid epoxidase in fosfomycin biosynthesis by a cation- and flavin-dependent mechanism. Proc Natl Acad Sci U S A. 2005 Oct 4;102(40):14221-6. Epub 2005 Sep 26. PMID:16186494
- ↑ Watanabe F, Yu F, Ohtaki A, Yamanaka Y, Noguchi K, Yohda M, Odaka M. Crystal structures of halohydrin hydrogen-halide-lyases from Corynebacterium sp. N-1074. Proteins. 2015 Dec;83(12):2230-9. doi: 10.1002/prot.24938. Epub 2015 Oct 16. PMID:26422370 doi:http://dx.doi.org/10.1002/prot.24938
- ↑ Yun D, Dey M, Higgins LJ, Yan F, Liu HW, Drennan CL. Structural Basis of Regiospecificity of a Mononuclear Iron Enzyme in Antibiotic Fosfomycin Biosynthesis. J Am Chem Soc. 2011 Jun 30. PMID:21682308 doi:10.1021/ja2025728