This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
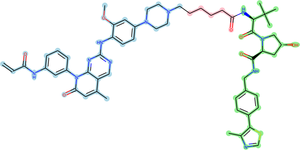
PROTAC
From Proteopedia
| |||||||||||
References
- ↑ Gadd MS, Testa A, Lucas X, Chan KH, Chen W, Lamont DJ, Zengerle M, Ciulli A. Structural basis of PROTAC cooperative recognition for selective protein degradation. Nat Chem Biol. 2017 Mar 13. doi: 10.1038/nchembio.2329. PMID:28288108 doi:http://dx.doi.org/10.1038/nchembio.2329
- ↑ PRosettaC https://prosettac.weizmann.ac.il, a computational resource for the prediction of PROTAC-induced ternary complexes.
- ↑ Zaidman D, Prilusky J, London N. PRosettaC: Rosetta based modeling of PROTAC mediated ternary complexes. J Chem Inf Model. 2020 Sep 25. doi: 10.1021/acs.jcim.0c00589. PMID:32976709 doi:http://dx.doi.org/10.1021/acs.jcim.0c00589
- ↑ PROTACpedia https://protacpedia.weizmann.ac.il/, a resource of manually curated data on Proteolysis Targeting Chimeras (PROTACs).