Introduction
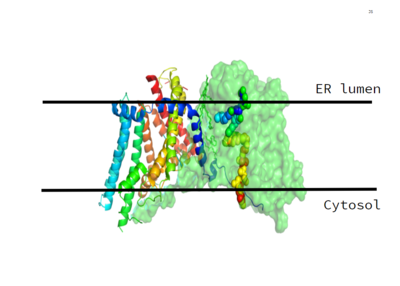
Figure 1. This image shows the location of the DGAT protein within the Endoplasmic Reticulum Membrane
DGAT, or Diacylglycerol Acyltransferase is a polytopic endoplasmic reticulum membrane protein embedded within the membrane of the ER. DGAT is highly expressed in epithelial cells of the small intenstine of homo sapiens. It can also be found in the liver, where it helps synthesize fats for storage, and the female mammary glands, where it produces fat in the milk.
DGAT was originally discovered by its homology to Acyl-CoA cholesterol acyltransferases (ACAT) 1 and 2. The structure, catalytic mechanism of diacylglycerol acyltransferase, and how DGAT interacts with CoA was discovered using a Cryo-EM. The Cryo-EM map revealed that DGAT forms a dimer, with each subunit containing nine transmembrane helices. The N and C terminals of each helice are located on the cytosolic and luminal sides of the endoplasmic reticulum membrane respectively.
Function
DGAT makes triglycerides from a diglyceride in plasma. In order to do this, DGAT uses two substrates: a fatty acyl-CoA and a diacylglycerol substrate. The basic mechanism consists of a lone pair on a hydroxyl group of glycerol attacking the carbon of the thioester bond of CoA. This results in the breakage of the thioester bond, and the attached acyl group attaches to the glycerol, creating a triglyceride.
Structure
DGAT consists of two domains, one cytoplasmic and one luminal. The cytoplasmic domain interacts with the interior of the cell and relays signals. The luminal domain senses misfolded proteins. The structure of DGAT consists of two protein chains, one ligand, two polymers, eighteen alpha helices and zero beta sheets. The majority of the transmembrane helices present within the structure form a concave-shaped ridge on either side of the membrane.
The DGAT dimer structure is formed primarily through many hydrogen-bonding interactions between the first 20 resolved residues (His69-Gly87). Hydrophobic interactions of the transmembrane helix region (Phe82-Ile98) with the other monomer also support the dimer structure formation. Additionally, there are four phospholipids present at the dimer interface that have been thought to contribute to the interactions between DGAT monomers.
Tunnels
Active Site
The active site of DGAT is located within the membrane, with the catalytic histidine residue (-represented in white) buried inside the central cavity. This central cavity serves as the catalytic site. The acyl-acceptor lipid substrates access the active site through the lateral gate within the membrane. The active site also contains (represented in white) and several nearby polar residues (including Asn378, Gln437, and Gln465) whose side chains are oriented towards the cavity center. These residues interact and create a hydrophilic channel within the active site. The His415 residue is also likely involved in catalysis, making it increasingly significant.
Relevance
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.

