Sandbox reserved 1751
From Proteopedia
Contents |
DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX
| |||||||||||
DNA Mismatch Repair by MutH
DNA Mismatch Repair (MMR) occurs when a mismatch of DNA bases occurs during DNA replication that is not corrected by the polymerases. This mismatch can be a at a single nucleotide or an insertion or deletion of up to 4 bases. An integral protein in MMR is MutH. MutH is an endonuclease, which means it is an enzyme that can digest DNA in the middle of the sequence. However, it is a weak endonuclease so it will only cause a nick upstream or downstream of the damaged daughter strand and allow it to be re-replicated by DNA polymerases. Homodimers of MutS and MutL bind the mismatched DNA and create a loop that MutH can bind to. In order to maintain the correct sequence and repair the damage without mutations, MutH must be able to differentiate the incorrect daughter strand from the correct parent strand. In bacteria, the freshly replicated DNA is hemimethylated, meaning that the parent strand is methylated and the daughter strand has not yet been methylated by methyltransferases. MutH then nicks the phosphodiester bond 5' of a GATC palindrome on the umethylated daughter strand. The GATC palindrome can be upstream or downstream of the damaged DNA site by up to 1000 nucleotides. This allows the damaged strand to be destroyed by exonucleases and re-replicated by DNA polymerase.
Structure of MutH
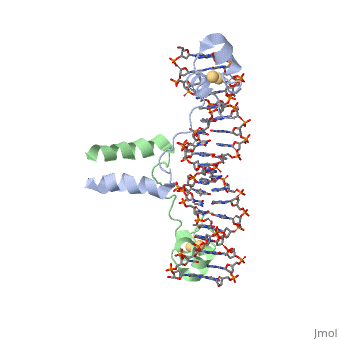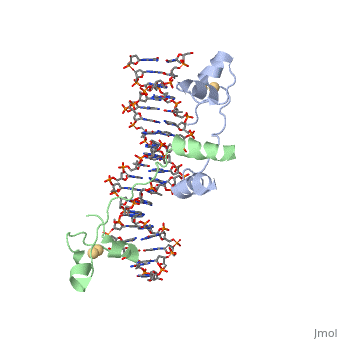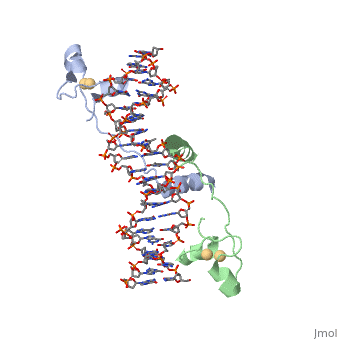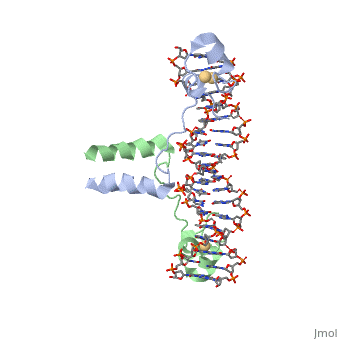
MutH has two subdomains, the "N" arm and the "C"arm. These arms are arranged in a "V" shape. The N arm contains the catalytic core, DEK motif and an essential Glu56 residue. The DEK motif consists of Asp-X(n)-Glu-X-Lys, and contains the Mg2+ required for nicking the phosphodiester bond. The DEK motif is found in most endonucleases. The C arm is responsible for base recognition and sequence-specific binding of the DNA. The cleft in the V binds the DNA. The C-term residues help to bind the N-arm and are shown to increase DNA binding in the closed position. The primary sequence of a protein provides it with its secondary and tertiary structure. This allows it to bind to the appropriate substrate and catalyze the reaction. This is the case for MutH as well when recognizing the GATC palindrome and nicking it correctly.
The Beta sheets 3/9/6 and loop 67 of arm "C" bind the GATC sequence in the major groove of the DNA. The N-arm contacts 6 nucleotides of the cleavage strand in the minor groove of the DNA. Loop C1S65 H-bonds nitrogen of Ala67 to stabilize the loop. Loop 67 (184-190) binds the GATC motif. G and C are hydrogen bonded by Asp184/Glu91 and Lys186/Gly187. Tyr212 bonds N6 the of unmodified adenine and Pro185 interacts with methylated adenine. These specific bonds allow for the recognition of hemimethylated DNA and differentiate the parent strand from the daughter strand. Loop BC Lys48 binds the O2s of the T’s. Also, Lys45/Asp46 interacts with the phosphate backbone to narrow the minor groove. The active site is supported to be Glu56, Asp70, Glu77, and Lys79, this makes up the DEK motif. The carboxylates (Glu/Asp) coordinate two Ca+ ions in the active site. Lys79, the 3’ phosphate of DNA, and the nearby metal ion activate water for a nucleophilic attack to create a single-stranded nick in the daughter strand 5' to the GATC palindrome. Lys79 links the two arms of MutH and allows for the sequence-specific cutting of DNA.
References
Ban, C., & Yang, W. (1998). Structural basis for MutH activation in E.coli mismatch repair and relationship of MutH to restriction endonucleases. The EMBO journal, 17(5), 1526–1534. https://doi.org/10.1093/emboj/17.5.1526
Lee, J. Y., Chang, J., Joseph, N., Ghirlando, R., Rao, D. N., & Yang, W. (2005). MutH complexed with hemi- and unmethylated DNAs: coupling base recognition and DNA cleavage. Molecular cell, 20(1), 155–166. https://doi.org/10.1016/j.molcel.2005.08.019
Voet, D., Voet, J. G., & Pratt, C. W. (2013). Fundamentals of Biochemistry: Life at the molecular level. Wiley.