This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox reserved 1752
From Proteopedia
Contents |
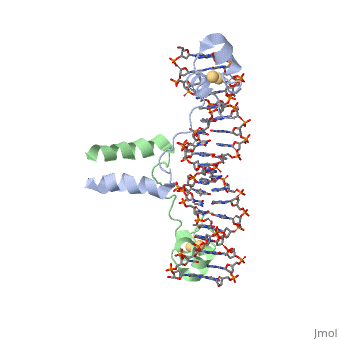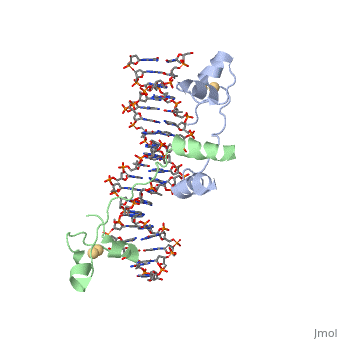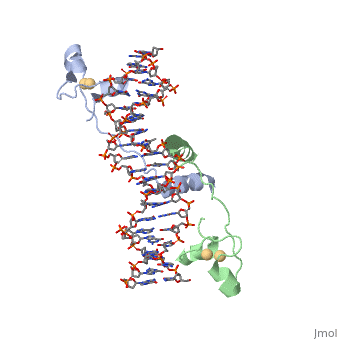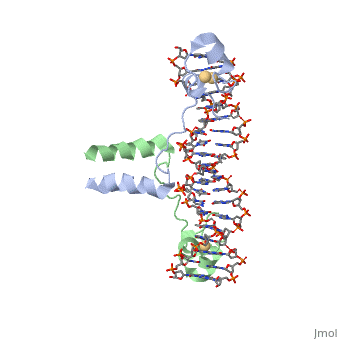
DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX
| |||||||||||
UvrD
|
, also known as Helicase II, is one of many components responsible in repairing DNA damage. Helicases use energy from nucleoside triphosphate hydrolysis to unwind double helices in metabolic pathways using nucleic acids. A nucleoside triphosphate (NTP) is a nucleotide with a nitrogenous base bound to a 5-carbon sugar with 3 phosphates attached. Phosphates are typically used to store energy that is released when breaking bonds to drive catabolic reactions. Helicases are also involved in multiple processes, including RNAs and DNAs, single stranded and double stranded. Helicases were found in the 1970’s to be DNA-dependent ATPases, meaning that they use ATP hydrolysis to complete its interactions with the different types of nucleic acids it comes into contact with. There are multiple types of helicases, with some being for RNA and DNA. Helicase II, also called UvrD is the founding member of SF1, one group of six superfamiliies used to identify helicases. SF1 and SF2 members share seven conserved sequence motifs that are involved in NTP binding. UvrD is important in replication, recombination, and repair from ultraviolet damage and mismatched base pairs. Go into NER from the book..
UvrD Motifs
There are 7 sequence motifs and a Q motif conserved among the SF1 and the SF2. I, Ia, II-VI are involved in ATP binding. Motifs Ia, III, and V are also involved in ssDNA binding. Motif IV is reported to be unique in SF1. They found in their paper, seven new sequence motifs conserved among UvrD homologs. They are Ib, Ic, Id, IVb, IVc, Va, and VIa. These conserved residues are involved in DNA binding or domain 1B and 2B interactions. In total, there are for UvrD, which are conserved in other homologous structures, such as PcrA, Rep, and Srs2. The homologous structures mentioned are Helicase 2 homologs, which appear in different species. These conserved motifs are important to maintain the function of UvrD.
UvrD Binding Site for ATP analog (AMPPNP)
When determining the structure of UvrD, an ATP analog was used. They used an so that the last phosphate can't be cleaved. Using the unhydrolyzable analog is beneficial in locking in the structure to observe.The green ion shown in the ATP analog scene is a Mg2+ ion, which is essential for ATP hydrolysis and interacts with the β and γ phosphates. The magnesium ion is surrounded by essential residues that when altered, have been shown to have reduced ATPase activity.
UvrD Binding Site for ATP analog (ADP*MgF3)
GIG Motif