This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox reserved 1752
From Proteopedia
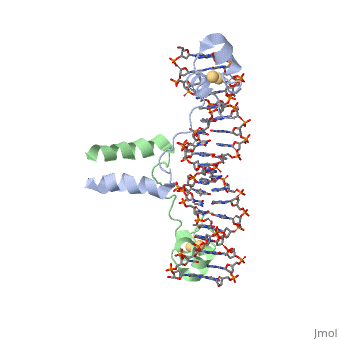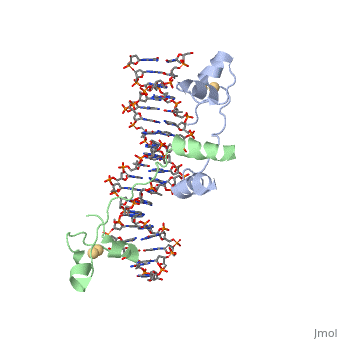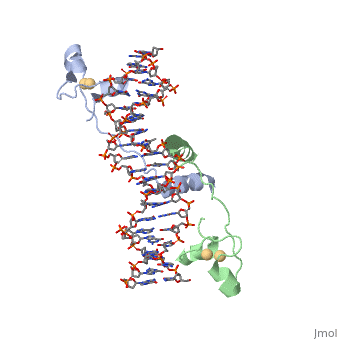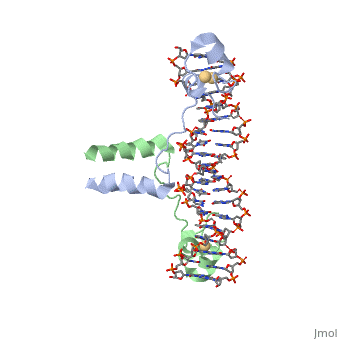
==DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX==
| |||||||||||
UvrD
, also known as Helicase II, is one of many components responsible in repairing DNA damage. Helicases use energy from ATP to unwind double helices in metabolic pathways using nucleic acids. ATP molecules are typically used to store energy shared between phosphate groups that gets released when breaking bonds to drive catabolic reactions.
Helicases were found in the 1970’s to be DNA-dependent ATPases, meaning that they use ATP hydrolysis to complete its interactions with the different types of nucleic acids it comes into contact with. Helicase II, also called UvrD is the founding member of SF1, one group of six superfamiliies used to identify helicases. SF1 and SF2 members share seven conserved sequence motifs that are involved in ATP binding RasMol [1] [2] Voet, Voet, & Pratt. (n.d.). Fundamentals biochemistry 4e : Free download, borrow, and streaming. Internet Archive. Retrieved October 11, 2022, from https://archive.org/details/FundamentalsBiochemistry4e