We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Journal:Proteins:3
From Proteopedia

Do Newly Born orphan proteins resemble Never Born proteins? A study using three deep learning algorithms
Newly Born proteins, or orphan proteins, have no sequence homology to other proteins and occur in single species or within a taxonomically restricted gene family
Never Born proteins are random polypeptides with amino acid content similar to that of native proteins.
Can recently developed AI/Deep Learning tools for predicting 3D protein structures like:
- AlphaFold2 (AF2)
- RoseTTAFold (RTF)
- Evolutionary Scale Modeling (ESM-2)
be useful to see if Newly Born proteins are similar to Never Born proteins?
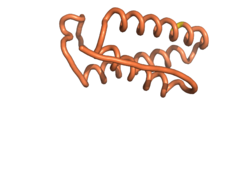
Sequences of Never Born proteins by Tretyachenko et al.[1] showed experimentally that some folded into compact structures, e.g., Never Born Protein 1856 is predicted to be compact for the 5 models predicted by RTF and AF2 and the single model by ESM-2 [Fig. 1].
| RTF-1856 | ESM-1856 | AF2-1856 | ||||||
|---|---|---|---|---|---|---|---|---|
| 
|
This page complements a publication in scientific journals and is one of the Proteopedia's Interactive 3D Complement pages. For aditional details please see I3DC.
