This is a default text for your page Ben Whiteside. Click above on edit this page to modify. Be careful with the < and > signs.
You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
Introduction
When consuming food, the human body is tasked with secreting hormones and chemical messengers that will help regulate homeostasis. After a meal, our body has to maintain homeostasis by reducing our blood glucose and signaling that we have consumed enough nutrients. During feeding, cells in the body will secrete the ligand, amylin. Amylin is a 37 amino acid glucoregulatory hormone that is produced within beta cells of the pancreas. When there is an influx of nutrients in the gastrointestinal tract, the ligand will bind to the heterodimeric receptor, activating the receptor and triggering the corresponding signaling cascade. The overall effect of this cascade is increased satiety, delayed gastric emptying, and inhibition of glucagon secretion. The amylin receptors are widely distributed throughout the central nervous system. The amylin g-protein coupled receptor is a heterodimeric protein containing a calcitonin receptor domain, as well as one of three receptor activity modifying proteins (RAMP 1,2, or 3).
[3]
[4]
[5]
Transmembrane Domain
Within the transmembrane domain (TMD) of the CTR, hydrophobic R groups span the phospholipid bilayer, anchoring the protein into the cell membrane upon amylin binding to the receptor.
Chemical Modifications to Amylin
N-Terminus Disulfide
The amylin peptide contains a between residues C2 and C7. This disulfide provides stability and rigidity to the helical structure of the peptide, allowing for favorable binding to the extracellular domain (ECD). Notable interactions formed by this disulfide include hydrogen bonds between E294 of the transmembrane domain with K1 of amylin, and both R362 and W361 of the transmembrane domain forming a hydrogen bond with N3 of amylin.
Amidated C-Terminus
The of amylin contains an amide group, rather than a carboxylic acid group. This chemical modification allows for more extensive hydrogen bonding to nearby residues, due to the added hydrogen bond donor on the NH2 group. In turn, this allows for favorable hydrogen bonds between S129 of the transmembrane domain and the main chain of Y37 on amylin. This interaction causes a "kink" in the random coil of amylin, displacing Y37 into a hydrophobic pocket, allowing for favorable hydrophobic interactions with W79 of the transmembrane domain. This amidation is thought to be a post-translational modification.
Two-Domain Model of Amylin Binding
It is hypothesized that amylin binds to the receptor via a two-domain model. The model suggests a series of steps for how amylin binds. First, the c-terminus of amylin binds to the n terminus of the extracellular domain of the receptor. This binding factors the alignment of amylin's n-terminus to the primary GPCR binding site. This activates the GPCR, leading to subsequent activation of adenylyl cyclase and cAMP release.
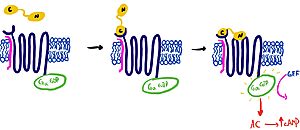
Figure 2: The Two Domain Model
RAMP-CTR Interface
is a key interaction that stabilizes the protein complex and positions the receptor to favorably bind to amylin. The RAMP-CTR interface extends into the plasma membrane, providing additional non-covalent bonding between the protein complex and the cell membrane.
Extracellular Domain - RAMP interactions
The extracellular domain of the CTR primarily contains polar residues in the extracellular space. In order to orient these residues in such a way to facilitate amylin binding, RAMP makes hydrogen bonds with the CTR to increase the rigidity of the receptor binding site.
Bypass Motif
G-alpha Interactions with CTR TMD
To transduce the signal across the cell membrane, the binding of amylin will induce a conformational change that allows for the CTR to make favorable interactions with the G alpha subunit. Two interactions shown activate the G-protein and propel downstream signaling.
Clincial Significance
Drug Development
Pramlintide is a synthetic analog of amylin that is commonly used in accordance with mealtime insulin to help treat type 1 and 2 diabetic patients. This drug binds to AMYR competitively, increasing the AMYR GPCR signaling. Increased action of the AMYR receptor has been shown to modestly lower HbA1c levels, which is often accompanied by weight loss (cite 5). Pramlintide binds with more affinity than amylin due to mutations from hydrophobic residues A29, S28, S29, and S37 to proline. The proline residues increase the rigidity of the ligand by creating unfavorable phi and psi angles, which improves the ability of the ligand to bind AMYR. Pramlintide treatment has also been shown to consistently reduce Amyloid β plaque aggregation in rodent models with Alzheimer’s disease (Gingell et al. 2014).
It has been thought that missense mutations (BLUE LINK) in residues C2 and C7 of the amylin peptide could lead to an increased risk of Alzheimer's Disease (CITE). Because of the rigidity these cysteine resides provide, reductions of their disulfide interaction leads to an increased risk of amyloid plaques. During drug design, pharmaceutical companies have focused on maintaining amylin residues, conserving C2 and C7, as well as K1, which forms a acts as a hydrogen bond donor for the E294 side chain and . Additionally, pharmaceuticals companies have also opted to maintain residues , which are critical residues in stabilizing the C terminus of amylin to the receptor binding site.
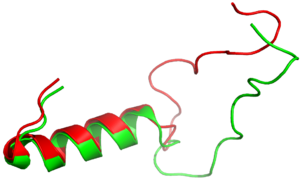

Figure 3:Amylin (green) aligned with Pramlintide (red)


