Electrostatic potential maps
From Proteopedia
It is revealing to visualize the distribution of electrostatic charges, electrostatic potential, on molecular van der Waals surfaces. Most protein-protein and protein-ligand interactions are largely electrostatic in nature, via hydrogen bonds and ionic interactions. Their strengths are modulated by the nature of the solvent: pure water or high ionic strength aqueous solution.
Contents |
Gallery
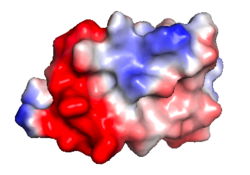
| Protein 1pgb is in the same orientation in all images. Positive + / Negative - | ||
|---|---|---|
| 
| 
|
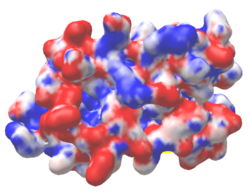
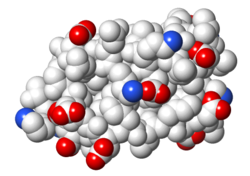
| Electrostatic potential map rendered by PyMOL using default molecular surface probe radius 1.4 Å. Method. | Electrostatic potential map rendered by iCn3D. | Van der Waals model colored by charge with FirstGlance in Jmol. Sidechain nitrogens on Arg/Lys; oxygens on Asp/Glu. |
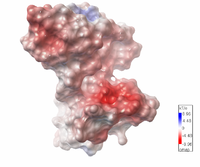
| Electrostatic potential map of 1tsj made with the Embedded Python Molecular Viewer from the Center for Computational Structural Biology of the Scripps Research Institute.
Click on the image to enlarge. |
Methods
===iCn3D
PyMOL
PyMOL has a license fee, but is free for students and educators.
- Download and install PyMOL.
- Enter command "fetch 1pgb".
- Menu: All, Action, remove waters.
- Menu: 1pgb, Action, generate, vacuum electrostatics, protein contact potential (local).
- Enter command "bg_color white".
Optional: The probe radius used to generate the molecular surface can be changed, and the previously generated surface will immediately change. The command is "set solvent_radius, 1.2" (don't overlook the comma!).
See Also
- Electrostatic interactions in Proteopedia.
- Jmol/Electrostatic potential methods.
- Isopotential Map in Wikipedia
- Delphi Web Server
