This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1a0i
From Proteopedia
|
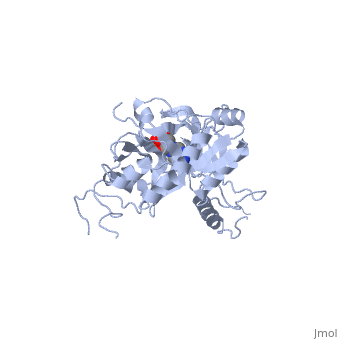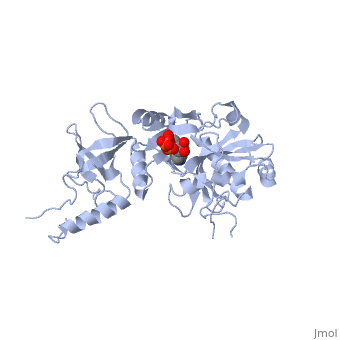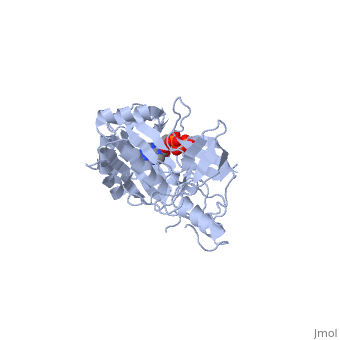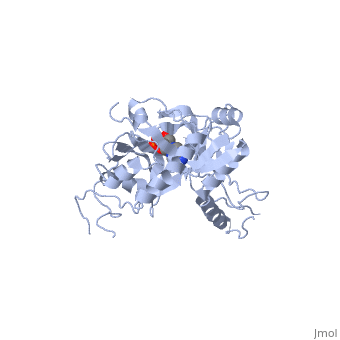
ATP-DEPENDENT DNA LIGASE FROM BACTERIOPHAGE T7 COMPLEX WITH ATP
Overview
The crystal structure of the ATP-dependent DNA ligase from bacteriophage, T7 has been solved at 2.6 A resolution. The protein comprises two domains, with a deep cleft running between them. The structure of a complex with, ATP reveals that the nucleotide binding pocket is situated on the larger, N-terminal domain, at the base of the cleft between the two domains of the, enzyme. Comparison of the overall domain structure with that of DNA, methyltransferases, coupled with other evidence, suggests that DNA also, binds in this cleft. Since this structure is the first of the, nucleotidyltransferase superfamily, which includes the eukaryotic mRNA, capping enzymes, the relationship between the structure of DNA ligase and, that of other nucleotidyltransferases is also discussed.
About this Structure
1A0I is a Single protein structure of sequence from Bacteriophage t7 with ATP as ligand. The following page contains interesting information on the relation of 1A0I with [DNA Ligase]. Active as DNA ligase (ATP), with EC number 6.5.1.1 Full crystallographic information is available from OCA.
Reference
Crystal structure of an ATP-dependent DNA ligase from bacteriophage T7., Subramanya HS, Doherty AJ, Ashford SR, Wigley DB, Cell. 1996 May 17;85(4):607-15. PMID:8653795
Page seeded by OCA on Sun Nov 18 08:56:34 2007